|
|
|
|
|
| Sample: |
Cyclohexanone monooxygenase monomer, 61 kDa Rhodococcus sp. HI-31 protein
|
| Buffer: |
50 mM Tris 5 mM NADP+, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Jul 15
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclohexanone monooxygenase monomer, 61 kDa Rhodococcus sp. HI-31 protein
|
| Buffer: |
50 mM Tris, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Jul 15
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclohexanone monooxygenase monomer, 61 kDa Rhodococcus sp. HI-31 protein
|
| Buffer: |
50 mM Tris 5 mM NADP+, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Jul 15
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
130 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclohexanone monooxygenase monomer, 61 kDa Rhodococcus sp. HI-31 protein
|
| Buffer: |
50 mM Tris 5 mM NADP+, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Jul 15
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclopentadecanone 1,2-monooxygenase monomer, 68 kDa Pseudomonas sp. HI-70 protein
|
| Buffer: |
50 mM Tris 2 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2013 Jan 27
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclopentadecanone 1,2-monooxygenase monomer, 68 kDa Pseudomonas sp. HI-70 protein
|
| Buffer: |
50 mM Tris 2 mM TCEP 5 mM NADP+, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Oct 2
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.7 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclopentadecanone 1,2-monooxygenase monomer, 68 kDa Pseudomonas sp. HI-70 protein
|
| Buffer: |
50mM Tris 2mM TCEP 5mM NADP+ 1mM cyclopentadecanon, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Oct 2
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclopentadecanone 1,2-monooxygenase monomer, 68 kDa Pseudomonas sp. HI-70 protein
|
| Buffer: |
50mM Tris mM TCEP 5mM NADP+ 1mM ω-pentadecalactone, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Oct 2
|
The role of conformational flexibility in Baeyer-Villiger monooxygenase catalysis and structure.
Biochim Biophys Acta 1864(12):1641-1648 (2016)
Yachnin BJ, Lau PCK, Berghuis AM
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
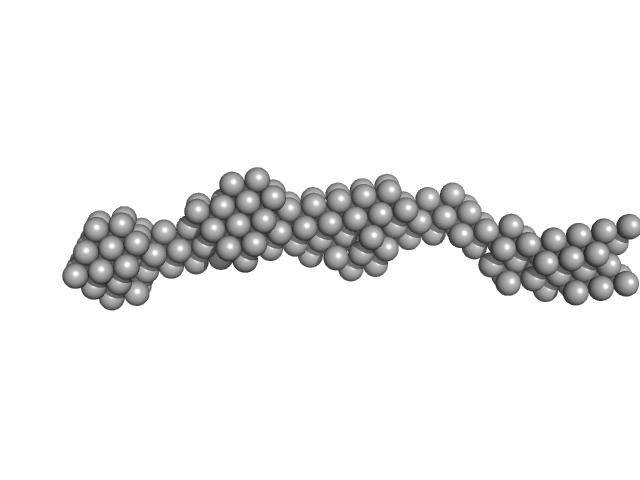
Bromodomain-containing protein 4 monomer, 56 kDa Homo sapiens protein
|
| Buffer: |
20mM Hepes, 100mM NaCl, 1mM Tris(2-carboxyethyl)phosphine hydrochloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 12
|
Potent and selective bivalent inhibitors of BET bromodomains.
Nat Chem Biol 12(12):1097-1104 (2016)
Waring MJ, Chen H, Rabow AA, Walker G, Bobby R, Boiko S, Bradbury RH, Callis R, Clark E, Dale I, Daniels DL, Dulak A, Flavell L, Holdgate G, Jowitt TA, Kikhney A, McAlister M, Méndez J, Ogg D, Patel J, Petteruti P, Robb GR, Robers MB, Saif S, Stratton N, Svergun DI, Wang W, Whittaker D, Wilson DM, Yao Y
|
| RgGuinier |
7.0 |
nm |
| Dmax |
27.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
Bromodomain-containing protein 4 monomer, 56 kDa Homo sapiens protein
|
| Buffer: |
20mM Hepes, 100mM NaCl, 1mM Tris(2-carboxyethyl)phosphine hydrochloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 12
|
Potent and selective bivalent inhibitors of BET bromodomains.
Nat Chem Biol 12(12):1097-1104 (2016)
Waring MJ, Chen H, Rabow AA, Walker G, Bobby R, Boiko S, Bradbury RH, Callis R, Clark E, Dale I, Daniels DL, Dulak A, Flavell L, Holdgate G, Jowitt TA, Kikhney A, McAlister M, Méndez J, Ogg D, Patel J, Petteruti P, Robb GR, Robers MB, Saif S, Stratton N, Svergun DI, Wang W, Whittaker D, Wilson DM, Yao Y
|
| RgGuinier |
6.2 |
nm |
| Dmax |
27.5 |
nm |
|
|