|
|
|
|
|
| Sample: |
Lipase chaperone monomer, 38 kDa Pseudomonas aeruginosa (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 100 µM TCEP, 5% glycerol,, pH: 8
|
| Experiment: |
SAXS
data collected at Xenocs Xeuss 2.0 Q-Xoom, Center for Structural Studies, Heinrich-Heine-University on 2019 Nov 18
|
Conformational dynamics of the membrane-anchored foldase LipH from Pseudomonas aeruginosa facilitates recognition and release of the client lipase
Jens Reiners
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
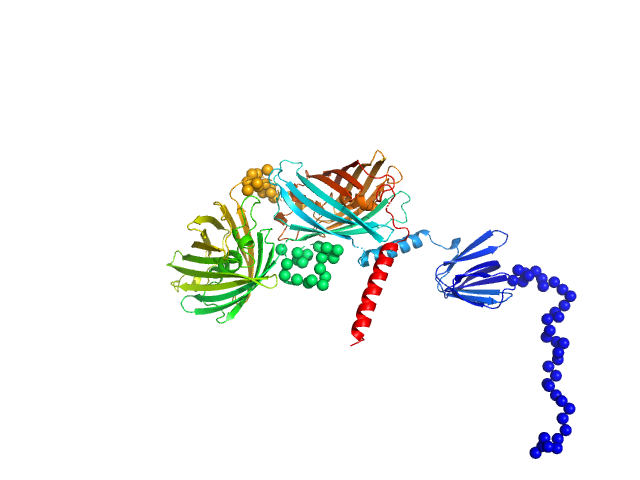
Ratiometric matryoshka multi-colour biosensor for ATP monomer, 72 kDa synthetic construct protein
|
| Buffer: |
50 mM MOPS, 100 mM KCl, 1 mM MgCl2,, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Nov 30
|
ATPLyzer – An advanced ratiometric multi-colour biosensor for long-term monitoring of ATP dynamics
Jens Reiners
|
| RgGuinier |
3.8 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
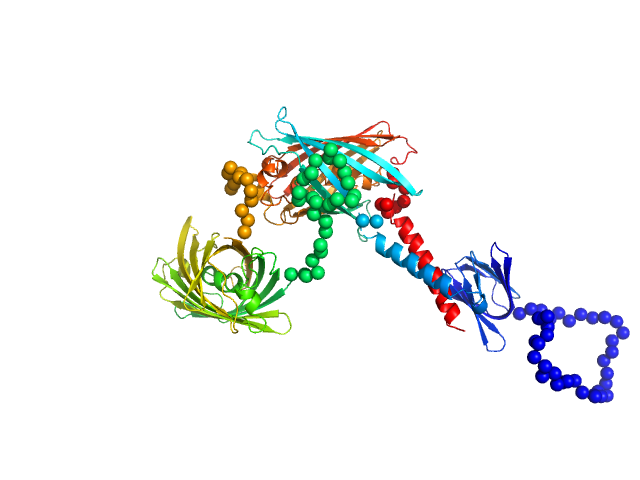
Ratiometric matryoshka multi-colour biosensor for ATP monomer, 72 kDa synthetic construct protein
|
| Buffer: |
50 mM MOPS, 100 mM KCl, 1 mM MgCl2, 5mM ATP,, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Nov 30
|
ATPLyzer – An advanced ratiometric multi-colour biosensor for long-term monitoring of ATP dynamics
Jens Reiners
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ubiquitin domain-containing protein DSK2 monomer, 39 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Feb 13
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
Ubiquitin domain-containing protein DSK2 monomer, 39 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Mar 7
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
Ubiquitin domain-containing protein DSK2 I45A monomer, 39 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Feb 13
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
DSK2 with HS1 deletion monomer, 37 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Feb 13
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
DSK2 with HS2 deletion monomer, 38 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Feb 13
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
DSK2 with HS3 deletion monomer, 38 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2026 Mar 7
|
Intramolecular Interactions between Folded and Disordered Regions Shape Ubiquilin Structure and Function
Advanced Science (2026)
Niblo J, Acharya N, Watkins M, Castañeda C, Sukenik S
|
|
|
|
|
|
|
|
| Sample: |
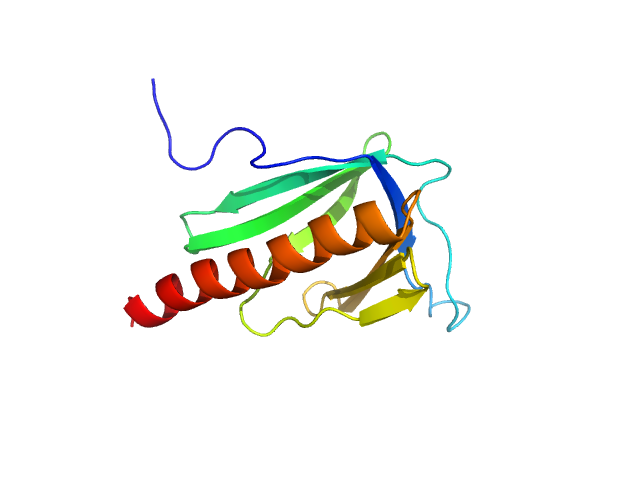
Point mutant M65I of EVH1 domain of Homer protein homolog 1 from mouse monomer monomer, 14 kDa Mus musculus protein
|
| Buffer: |
20mM NaCl, 50 mM NaPi, 0.02% of sodium-azide, pH: 7.4
|
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, CEITEC on 2023 Nov 8
|
Modulation of Homer1 EVH1 domain internal dynamics by putative autism-associated mutations.
FEBS Lett (2026)
Farkas F, Maruzs B, Kálmán ZE, Klumpler T, Batta G, Péterfia B, Gáspári Z
|
| RgGuinier |
1.7 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
24 |
nm3 |
|
|