Small Angle Scattering Biological Data Bank
Thanks to recent advances in instrumentation and computational methods, the amount of experimental SAS data and subsequent publications is increasing dramatically. The urgent need for a global repository that would allow investigators to locate and access experimental scattering data and derived models was stressed by the wwPDB small angle scattering task force (SAStf)1.
The Small Angle Scattering Biological Data Bank (SASBDB)2, 3 was developed in accordance with the plans of the SAStf, which foresee a development of a federated system of interconnected databases for SAXS/SANS. SASBDB is a curated repository of freely accessible and fully searchable SAS experimental data, which are deposited together with the relevant experimental conditions, sample details, instrument characteristic and derived models. The quality of deposited experimental data and the accuracy of models obtained from SAS and complementary techniques is assessed by the site developers. Following the SAStf recommendations, SASBDB consents to import and export data using sasCIF, an extension of core Crystallographic Information File for SAS4.
Most of the entries are published data and models from the studies where SAS was employed for the structural analysis of macromolecular solutions. There are also "benchmark" experimental data available from a set of well-characterized commercially available proteins. The SAXS data were collected with on-line purification, which ensures sample monodispersity. The high resolution structures of the benchmark set are available, and these data can be used for e.g. to test computational approaches for tutorials etc.
The data and models deposited in SASBDB are manually curated. Please sign in with your SASBDB online account if you wish to deposit your SAS data to SASBDB.
The data and models deposited in SASBDB are free of all copyright restrictions and made fully and freely available for both non-commercial and commercial use. Users of the data should attribute the original authors.
Scientific publications referring to SASBDB: https://europepmc.org/search?query=sasbdb
The use of SASBDB for methods development
Publications reporting the use of SASBDB for methods development are listed here.
Small angle scattering (SAS)
Small angle X-ray and neutron scattering (SAXS and SANS) are important research tools in structural biology. They allow us to investigate the three-dimensional structures and structural changes of biological molecules.
Small-angle scattering (SAS) experiences a renaissance in the studies of macromolecular solutions allowing one to study the structure of native particles and complexes and to rapidly analyse structural changes in response to variations in external conditions. Novel data analysis methods significantly enhanced resolution and reliability of structural models provided by the technique5. Emerging automation of the experiment, data processing and interpretation make solution SAS, especially synchrotron X-ray scattering (SAXS) on high brilliance synchrotron sources a streamlined tool for large scale structural studies in molecular biology. The method provides low resolution macromolecular shapes ab initio and is readily combined with other structural and biochemical techniques in hybrid approaches. Of special interest is the joint use of SAS with the high resolution methods like crystallography and NMR, but also with complementary biophysical and biochemical techniques. Rapid validation of predicted or experimentally obtained high resolution models in solution, identification of biologically active oligomers and addition of missing fragments to high resolution crystallographic models are possible. For macromolecular complexes, quaternary structure can be effectively analysed in terms of rigid body movements/rotations of individual subunits, if their structures are available. SAS can also be applied to characterize the solution states of flexible and intrinsically disordered macromolecules.
Small angle scattering experiment
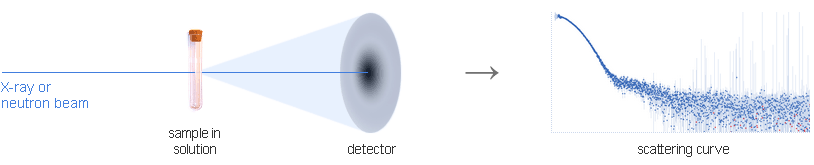
Conceptually, a small angle X-ray scattering experiment is simple: a solution of biological macromolecules is illuminated by X-rays and the scattered radiation is registered by a detector. Since the molecules in solution are randomly oriented, the 2D scattering pattern is isotropic and can be radially averaged into a 1D scattering curve that reflects how the scattered intensity changes with the angle. A small angle neutron scattering experiment is similar; neutrons are used instead of X-rays. SANS is preferable for studies of macromolecular complexes.
Statistics
SASBDB policies
Please read our SASBDB policies carefully.
References
- Trewhella J, Hendrickson WA, Kleywegt GJ, Sali A, Sato M, Schwede T, Svergun DI, Tainer JA, Westbrook J, Berman HM Report of the wwPDB Small-Angle Scattering Task Force: Data Requirements for Biomolecular Modeling and the PDB Structure. Structure 21:875-881 (2013) Europe PMC
- Valentini E, Kikhney AG, Previtali G, Jeffries CM, Svergun DI SASBDB, a repository for biological small-angle scattering data. Nucleic Acids Research 43(D1):D357–D363 (28 Jan 2015) Europe PMC
- Kikhney AG, Borges CR, Molodenskiy DS, Jeffries CM, Svergun DI SASBDB: Towards an automatically curated andvalidated repository for biological scattering data. Protein Science 29(1):66-75 Europe PMC
- M. Kachala, J. Westbrook and D. Svergun Extension of the sasCIF format and its applications for data processing and deposition. J. Appl. Cryst. 49:302-310 (2016)
- Svergun DI, Koch MHL, Timmins PA, May RP. Small Angle X-Ray and Neutron Scattering from Solutions of Biological Macromolecules. IUCr Texts on Crystallography, No. 19. Oxford University Press (2013)
Related resources
- PED - Protein Ensemble Database
- AlphaFold - Protein Structure Database
- wwPDB - Protein Data Bank
- EMDB - Electron Microscopy Data Bank
- BMRB - Biological Magnetic Resonance Bank
Contact us
SASBDB is powered by the Biological Small Angle Scattering Group, European Molecular
Biology Laboratory, Hamburg Outstation.
For questions and feedback please contact:
.
