| MWI(0) | 622 | kDa |
| MWexpected | 606 | kDa |
|
log I(s)
2.51×103
2.51×102
2.51×101
2.51×100
|
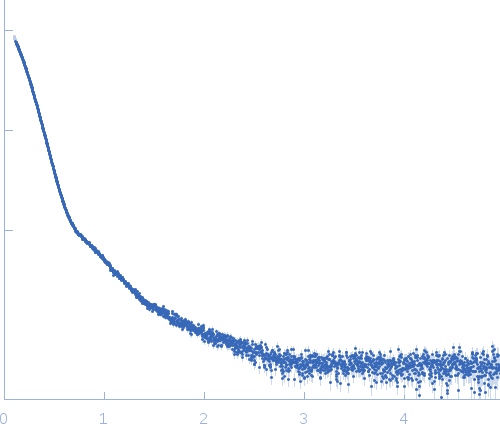 s, nm-1
s, nm-1
|
|
|
|

|
|
Synchrotron SAXS
data from solutions of
Thyroglobulin in Tris/HCl
in
100 mM Tris/HCl 100 mM NaCl, pH 7.5
were collected
on the
EMBL X33 beam line
at the DORIS III, DESY storage ring
(Hamburg, Germany)
using a MAR 345 Image Plate detector
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
One solute concentration of 3.90 mg/ml was measured
at 20°C.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
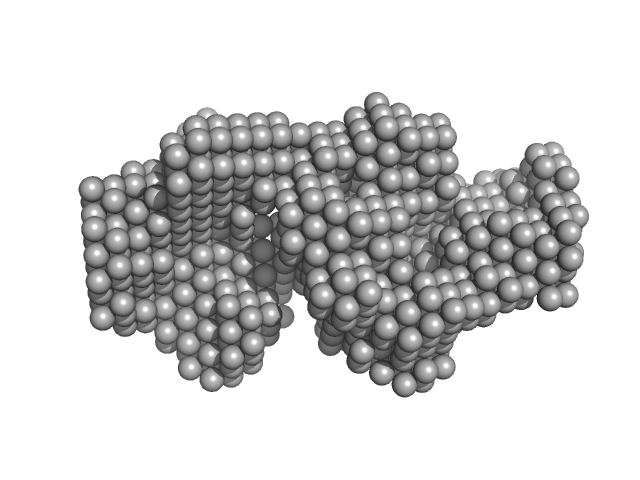
Thyroglobulin contains glycans and extra iodines, for these reasons the molecular weight based on the sequence (606 kDa) does not agree with the experimentally calculated one (622 kDa).
Tags:
X33
|
|
|||||||||||||||||||||||||||