|
Synchrotron SAXS data from solutions of heterohexameric CcdB2-CcdA2-CcdB2 toxin/antitoxin complex in 10 mM Tris 50 mM NaCl, pH 7.3 were collected using size exclusion chromatography SAXS (SEC-SAXS) on the SWING beam line at SOLEIL (Saint-Aubin, France) equipped with a CCD AVIEX detector at a sample-detector distance of 1.8 m and at a wavelength of λ = 0.103 nm (I(s) vs s, where s = 4πsinθ/λ and 2θ is the scattering angle). Data were collected in HPLC mode using a Shodex KW402.5-4F column that had been pre-equilibrated with running buffer for at least one column volume. Experiments were performed at 15°C using a flow rate of 0.2 ml/min. Prior to sample injection, the CcdB2-CcdA2-CcdB2 complex was prepared to a final concentration of 7 mg/ml by slowly titrating CcdA2 into an excess of CcdB2. The sample was then injected onto the column and 13 successive 0.750 second SAXS data frames were collected through the elution peak of the complex. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted and the different curves scaled and averaged to produce the scattering profile displayed in this entry.
Concentration min = UNKNOWN. Concentration max = UNKNOWN
|
|
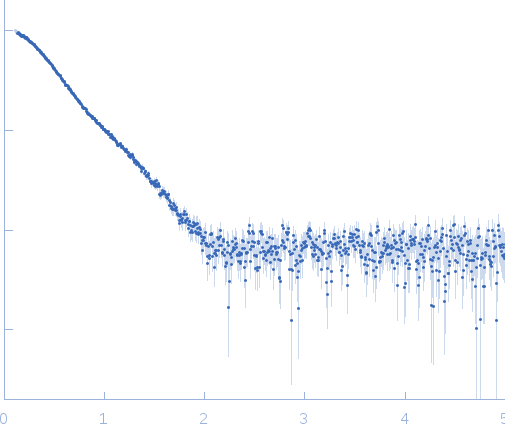 s, nm-1
s, nm-1