|
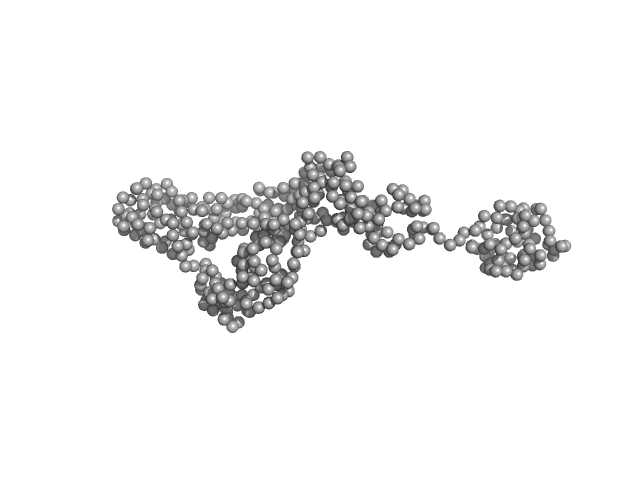
Synchrotron SAXS
data from solutions of
Human NEI like DNA glycosylase 1 (NEIL1)
in
25mM HEPES 300mM NaCl 1mM DTT 10% Glycerol, pH 7.5
were collected
on the
BioCAT 18ID beam line
at the Advanced Photon Source (APS), Argonne National Laboratory storage ring
(Lemont, IL, USA)
using a MAR 165 CCD detector
at a wavelength of λ = 0.1033 nm
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
Solute concentrations ranging between 0.5 and 2 mg/ml were measured
at 10°C.
14 successive
1 second frames were collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
Experimental MW reported here is calculated from Gnom based I(0) estimates and will be used for publication purpose in future. Guinier I(0) estimates are prone to presence of aggregates/radiation damage and that is why Gnom based I(0) was preferred to calculate the experimental MW.
|
|
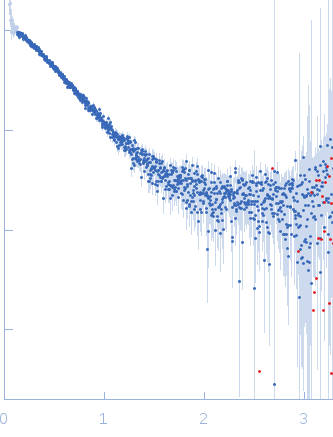 s, nm-1
s, nm-1