|
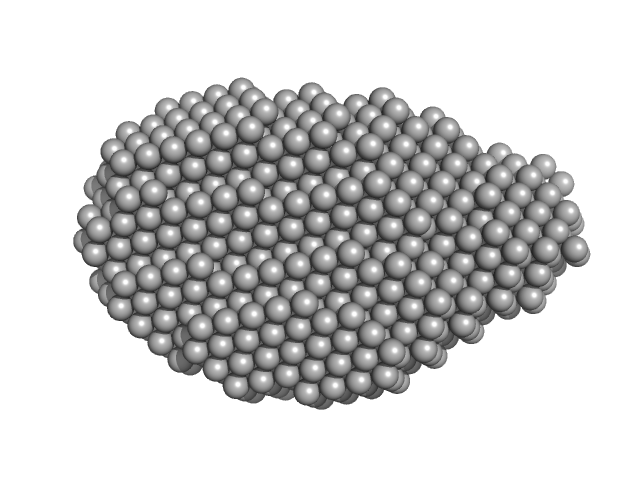
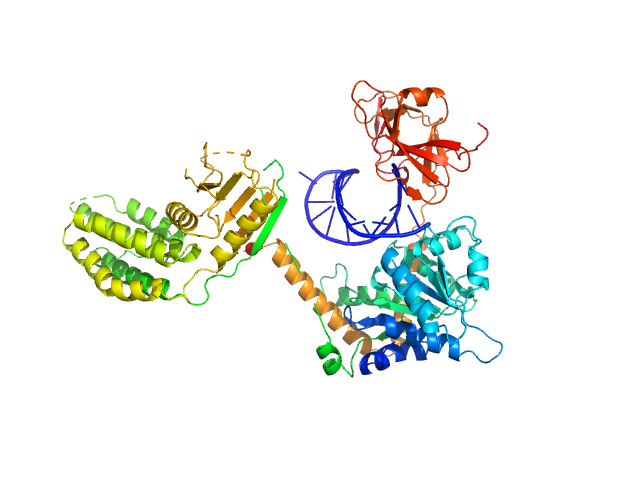
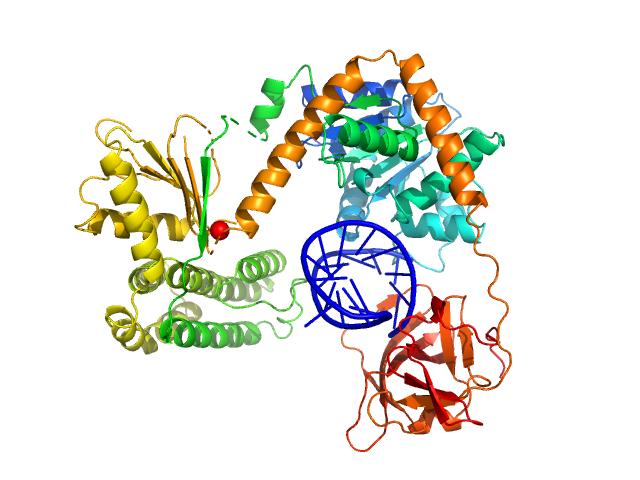
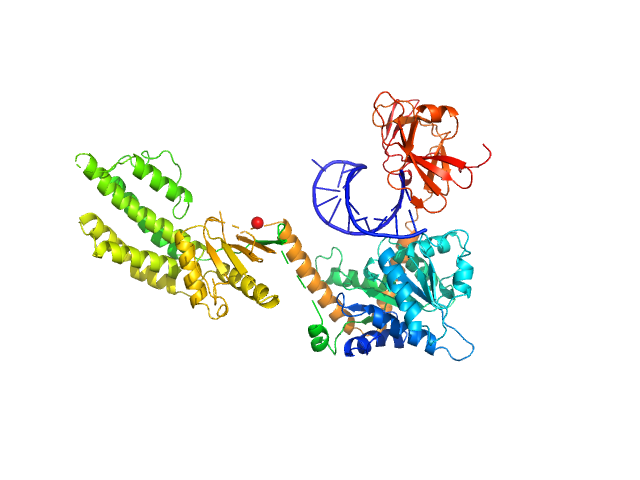
Synchrotron SAXS data from solutions of Probable ATP-dependent RNA helicase DDX58 (Delta-CARDs) plus 8mer hairpin dsRNA/AMP-PNP (SEC-peak1) in 25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, 0.5 mM AMP-PNP, pH 7.4 were collected using size exclusion chromatography (SEC-SAXS) on the SAXS/WAXS camera on the storage ring Australian Synchrotron (Melbourne, Australia) using a Pilatus 1M detector at a sample-detector distance of 1.6 m and at a wavelength of λ = 0.10332 nm (I(s) vs s, where s = 4π sin θ/λ and 2θ is the scattering angle). Data frames (11 x 5 s exposures) were collected through the SEC elution peak at 35.1 min (GE Healthcare Life Sciences Superdex 200 10/300; 0.4 ml/min; 50 μl injection at 8 mg/ml). The data were normalized to the intensity of the transmitted beam and radially averaged; the radially averaged scattering of the solvent-blank was subtracted and the different curves were scaled to yield the final composite scattering curve. The SAXS data were collected at 15 °C.
Concentration min = UNKNOWN. Concentration max = UNKNOWN
|
|
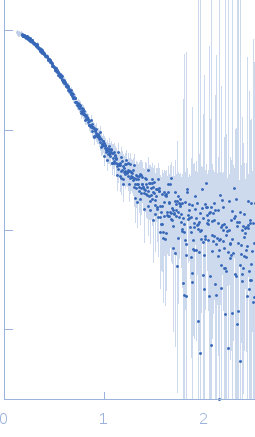 s, nm-1
s, nm-1