|
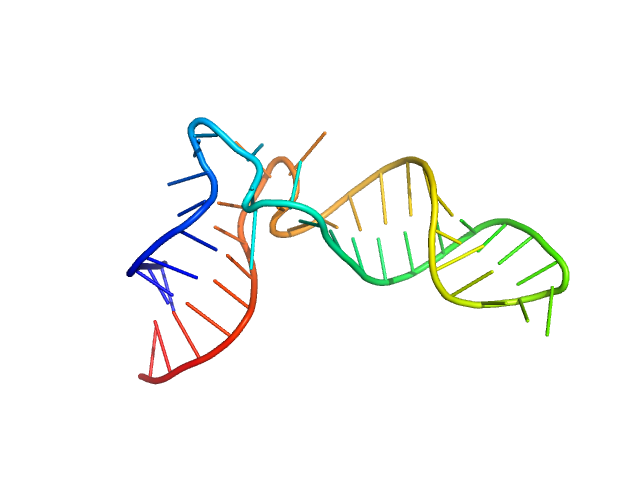
Synchrotron SAXS data from solutions of mitochondrial tRNA-Ile were performed at the SWING beam line (SOLEIL synchrotron, Saint-Aubin, France) equipped with a 17 x 17 cm2 low-noise Aviex CCD detector at a sample-detector distance of 2.8 m using an X-ray wavelength of λ = 0.1033 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). The tRNA-Ile sample was concentrated to 3.5 mg/ml in 200 µl buffer containing 50 mM HEPES-NaOH, pH 7.4, and 10 mM MgCl2 using centrifugation in an Amicon Ultra-4 (cut off 3 kDa). 70 µl of solution were loaded onto a size exclusion chromatography (SEC) HPLC column (BioSEC3-150, Agilent) upstream the SAXS cell in order to separate monomers from potential oligomers and aggregates. SEC elution was performed using an Agilent HPLC system at 0.2 ml/min in 50 mM HEPES-NaOH, pH 7.4, and 10 mM MgCl2 at 15°C. Samples were exposed to X-rays directly downstream to the SEC column to record the corresponding scattering signal (240 images of 1 sec exposure at 0.5 sec interval). The scattering signal from the buffer was collected at the beginning of the run (90 images) to be subtracted from the RNA signal. SAXS images were processed with FOXTROT (David & Pérez, 2009) to generate individual curves (subtracted from the buffer signal) and plots corresponding to I(0) and gyration radius Rg as a function of frames. US-SOMO (Brookes et al, 2016) was used to separate the scattering signal corresponding of successive molecular species in overlapping regions of the chromatogram by Gaussian decomposition and curves from consecutive images showing similar Rg were averaged. Resulting SAXS profiles were analyzed with the package ATSAS 2.8.2 (Franke et al, 2017) to evaluate structural parameters such as radius of gyration Rg from Guinier plots (PRIMUS; Konarev et al, 2003), as well as the particle distance distribution function p(r) (GNOM; Svergun, 1992) and associated maximum distance Dmax and Porod volume.
SEC column = UNKNOWN. Sample injection volume = UNKNOWN. Flow rate = UNKNOWN
|
|
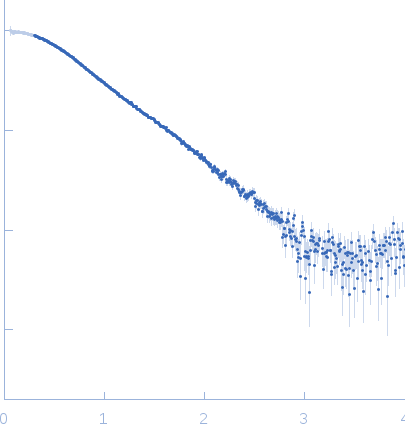 s, nm-1
s, nm-1