|
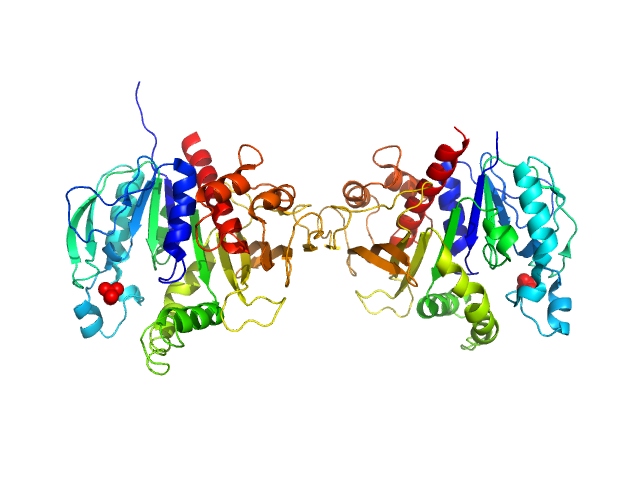
Synchrotron SAXS data from solutions of chloroplastic phosphoribulokinase (collected using SEC-SAXS) in Tris-HCl 50 mM 150 mM KCl, pH 7.5 were collected on the BM29 beam line at the ESRF (Grenoble, France) using a Pilatus 1M detector at a sample-detector distance of 2.9 m and at a wavelength of λ = 0.099 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). SEC-SAXS was performed at 4°C. 80 successive 1 second frames were collected through the SEC-elution profile. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of an appropriate solvent-blank (eluate SEC-buffer) was subtracted.
Additional SEC parameters: Column type: Superdex 200 10/300 GL (GE Healthcare); Flow rate: 0.5 ml/min; Sample injection concentration: 6.1 mg/ml; Injection volume: 100 µl.
|
|
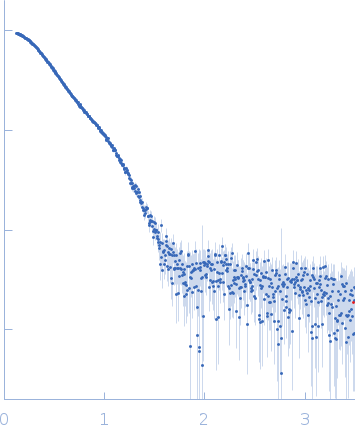 s, nm-1
s, nm-1