|
Synchrotron SAXS
data from solutions of
Unlabeled Nuclear Localization Signal (NLS) from the inner nuclear membrane protein HEH2 without denaturant
in
25 mM HEPES, 150 mM NaCl, 10 mM DTT, pH 7.4
were collected
on the
EMBL P12 beam line
at the PETRA III storage ring
(DESY; Hamburg, Germany)
using a Pilatus 2M detector
at a sample-detector distance of 3 m and
at a wavelength of λ = 0.1 nm
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
Solute concentrations ranging between 2 and 7.5 mg/ml were measured
at 23°C.
20 successive
0.050 second frames were collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
The low angle data collected at lower concentration were merged with the highest concentration high angle data to yield the final composite scattering curve.
The protein contains a penultimate non-canonical amino acid p-acetylphenylalanine (207 Da) that is represented as U (selenocysteine, 168 Da) in the amino acid sequence for the entry. Therefore, the calculated MW from sequence (MW(expected)) must be adjusted accordingly (ca. 40 Da).
|
|
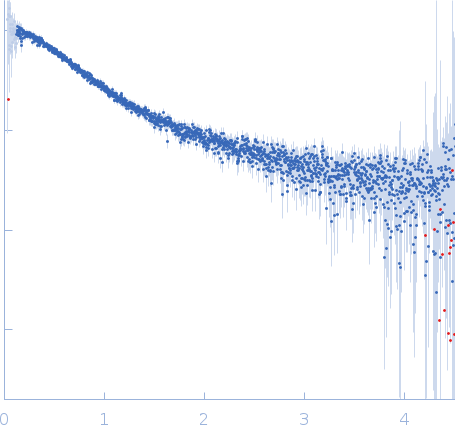 s, nm-1
s, nm-1
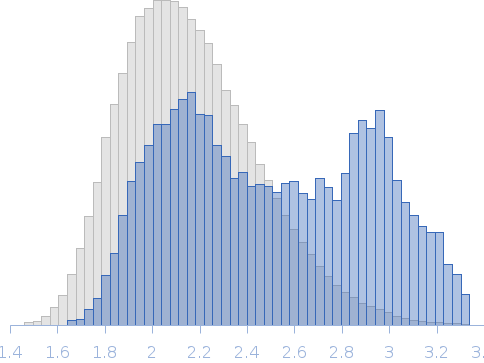 Rg, nm
Rg, nm