|
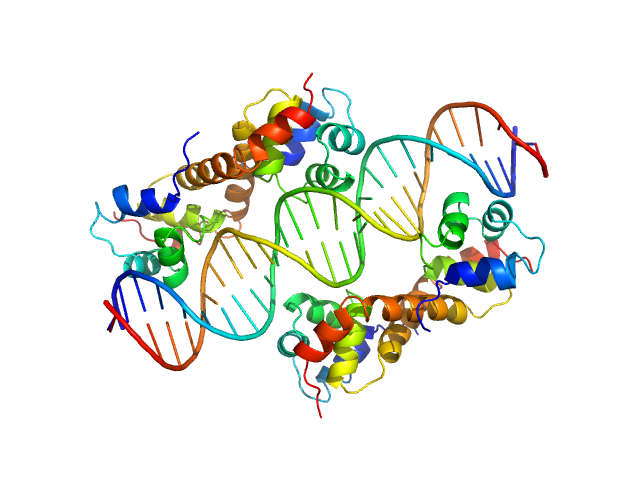
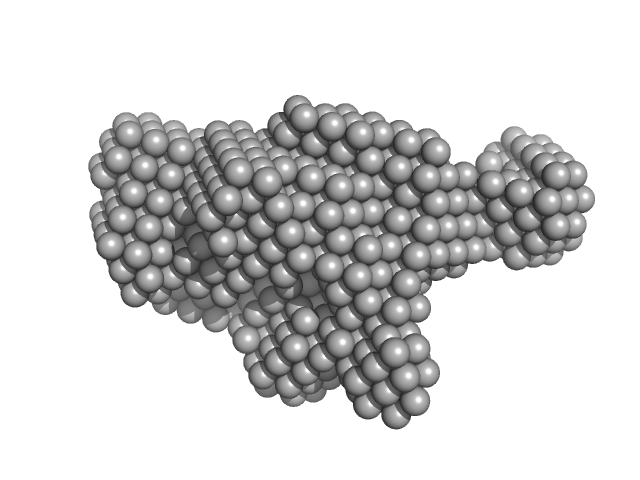
Synchrotron SAXS data from solutions of the PaHigA-DNA complex in 20 mM Tris, 300 mM NaCl, 5% (v/v) glycerol, 1 mM PMSF, pH 8 were collected on the BL19U2 beam line at the Shanghai Synchrotron Radiation Facility (SSRF; Shanghai, China) using a Pilatus 1M detector at a sample-detector distance of 2.7 m and at a wavelength of λ = 0.103 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 1 and 5 mg/ml were measured at 23°C. 20 successive 1 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The low angle data collected at lower concentration were merged with the highest concentration high angle data to yield the final composite scattering curve.
|
|
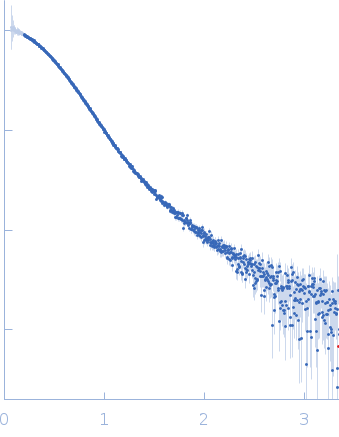 s, nm-1
s, nm-1