|
Synchrotron SAXS data from solutions of Ile-Leu-Glu-Ile-Asn-Ser (ILQINS) hexapeptide self-assemblies suspended in pure (MQ, 18 MΩ) water, pH 7, were collected on the SAXS/WAXS beam line at the Australian Synchrotron (Melbourne, Australia) using a Pilatus 1M detector at a sample-detector distance of 1.6 m and at a wavelength of λ = 0.103 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 1 and 5 mg/ml were measured at 25°C. 15 successive 1 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The low angle data collected at lower concentration were merged with the highest concentration high angle data to yield the final composite scattering curve. The errors on I(s) have not been propagated.
The SAXS/WAXS profile shows a solution of ILQINS peptide measured during the process of aggregation. The beam width is smaller than estimated size of (some) aggregates, so the calculation of Rg or of the self-assembled polymer molecular weight is not valid; the assemblies of the 0.7 kDa ILQINS peptide units are in the order of several hundred(s)/thousands of kDa. Related SASBDB entry: SASDHK8.
|
|
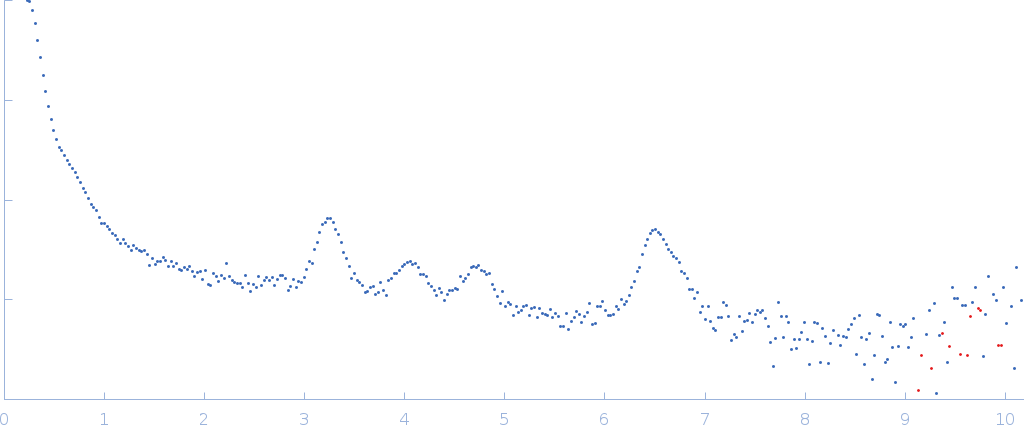 s, nm-1
s, nm-1
