|
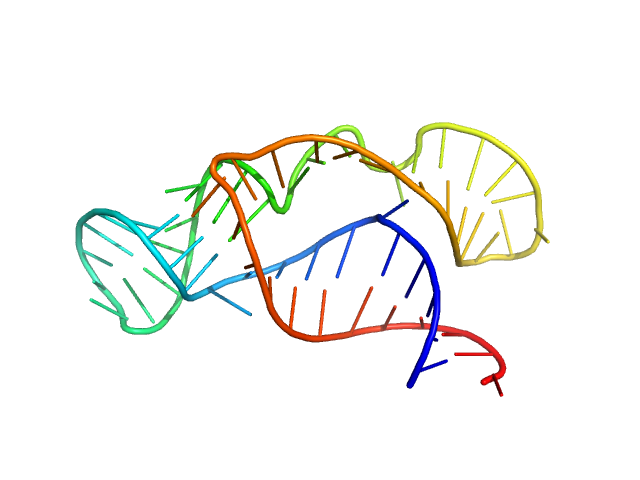
Synchrotron SAXS data from solutions of II-III-VI three-way junction from the VS ribozyme in 50 mM MES, 50 mM KCl, 5 mM MgCl2, pH 6.5 were collected on the G1 beam line at the Cornell High Energy Synchrotron Source (CHESS; Ithaca, NY, USA) using a Pilatus 100K detector at a sample-detector distance of 1.5 m and at a wavelength of λ = 0.1245 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 0.5 and 2 mg/ml were measured at 20°C. 10 successive 2 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The low angle data collected at lower concentration were merged with the highest concentration high angle data to yield the final composite scattering curve.
|
|
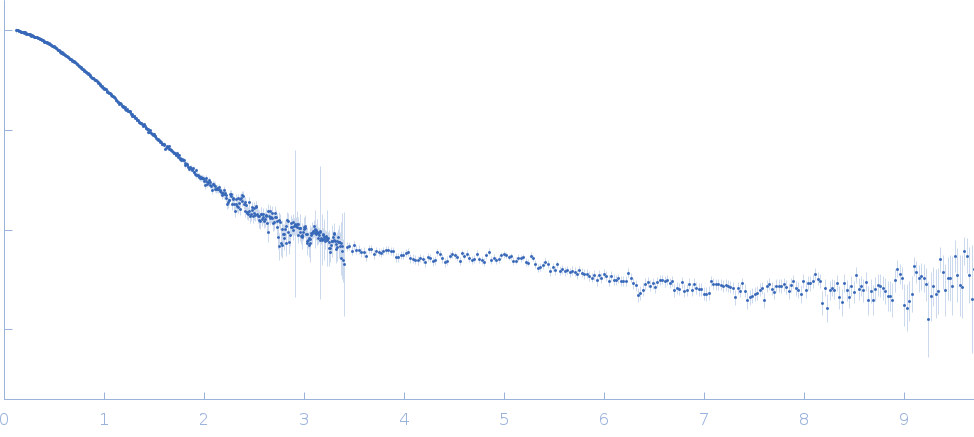 s, nm-1
s, nm-1