|
Synchrotron SAXS
data from solutions of
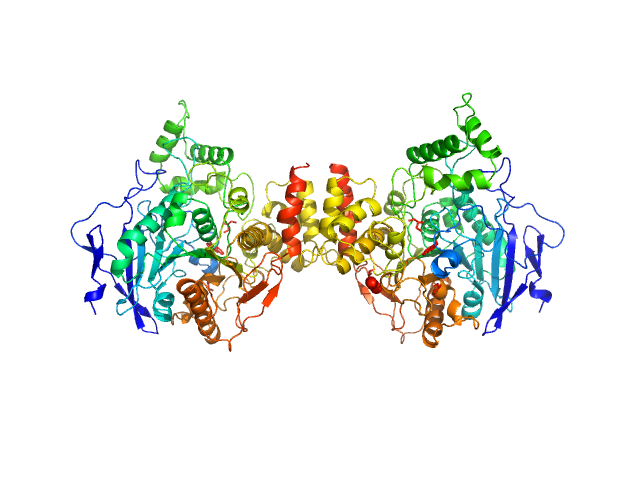
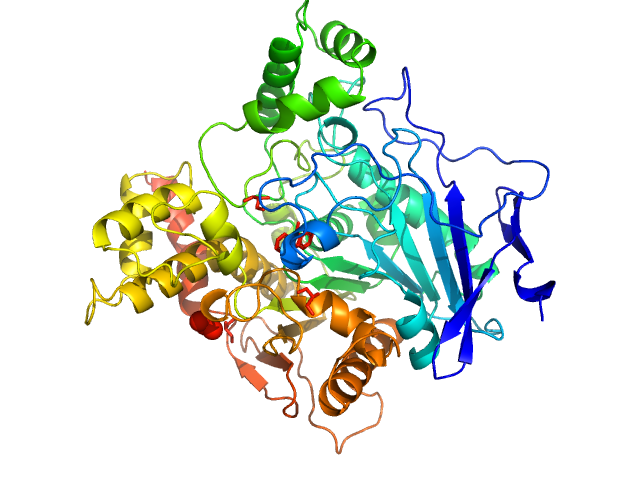
Human acetylcholinesterase, covalently bound to paraoxon
in
50 mM Tris/HCl, 100 mM NaCl, pH 7.4
were collected
on the
BL4-2 beam line
at the Stanford Synchrotron Radiation Lightsource (SSRL) storage ring
(Menlo Park, CA, USA)
using a Rayonix MX225-HE detector
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
One solute concentration of 2.00 mg/ml was measured
at 22°C.
10 successive
1 second frames were collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
The estimated experimental MW and shape of the PDDF indicate this sample represents a mixture of homodimer and monomer. X-ray wavelength: UNKNOWN.
|
|
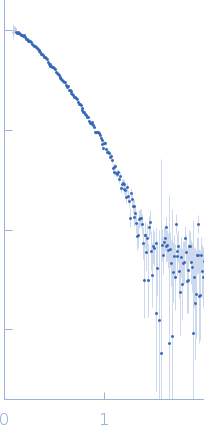 s, nm-1
s, nm-1