| MWexperimental | 14 | kDa |
| MWexpected | 12 | kDa |
| VPorod | 22 | nm3 |
|
log I(s)
1.92×101
1.92×100
1.92×10-1
1.92×10-2
|
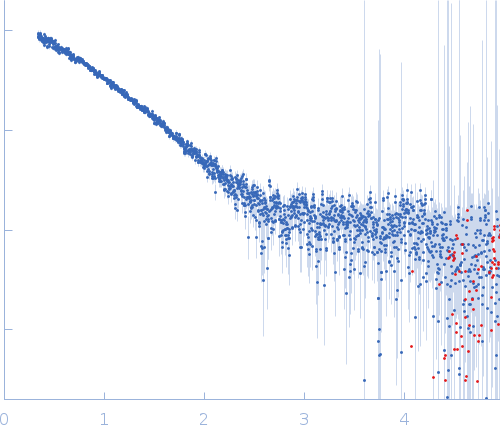 s, nm-1
s, nm-1
|
|
|
|
|
|

|
|
Synchrotron SAXS
data from solutions of
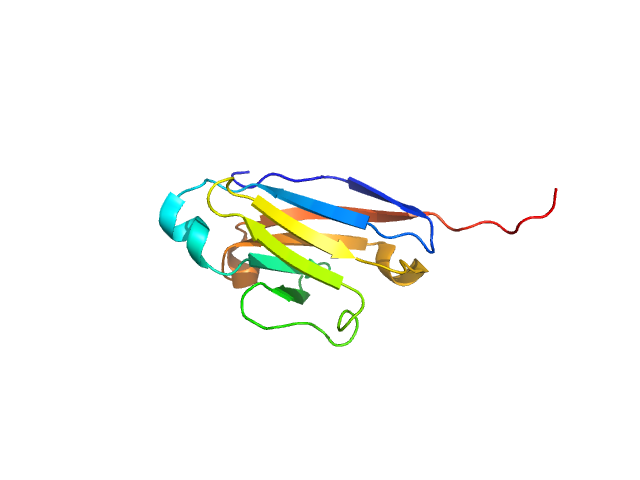
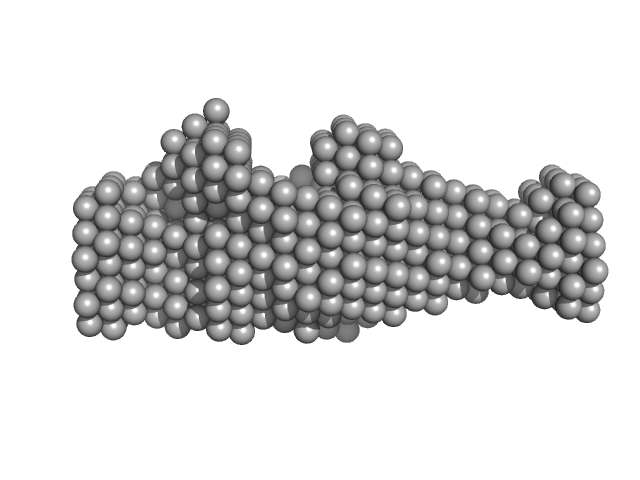
Human NK inhibitory receptor IRp60 with an immunoglobulin-like fold
in
MES buffer with 3mM DTT, pH 5.5
were collected
on the
EMBL X33 beam line
at the DORIS III, DESY storage ring
(Hamburg, Germany)
using a MAR 345 Image Plate detector
at a sample-detector distance of 2.4 m and
at a wavelength of λ = 0.15 nm
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
One solute concentration of 5.50 mg/ml was measured
at 8°C.
One
180 second frame was collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
Storage temperature = UNKNOWN
Tags:
X33
|
|
|||||||||||||||||||||||||||||||||