|
Synchrotron SAXS data from solutions of aconitate hydratase B, monomer fraction in 100 mM NaCl, 20 mM Tris, 1 mM DTT, 3% v/v glycerol, pH 8 were collected on the EMBL P12 beam line at PETRA III (DESY; Hamburg, Germany) using a Pilatus 6M detector at a sample-detector distance of 3 m and at a wavelength of λ = 0.124 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). One solute concentration of 4.90 mg/ml was measured at 20°C. 60 successive 0.100 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
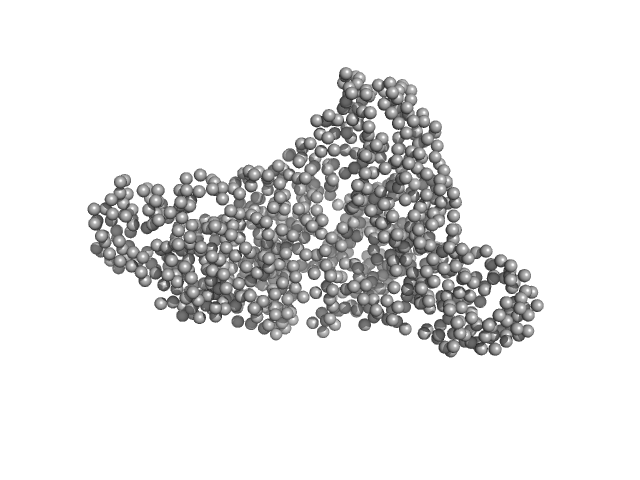
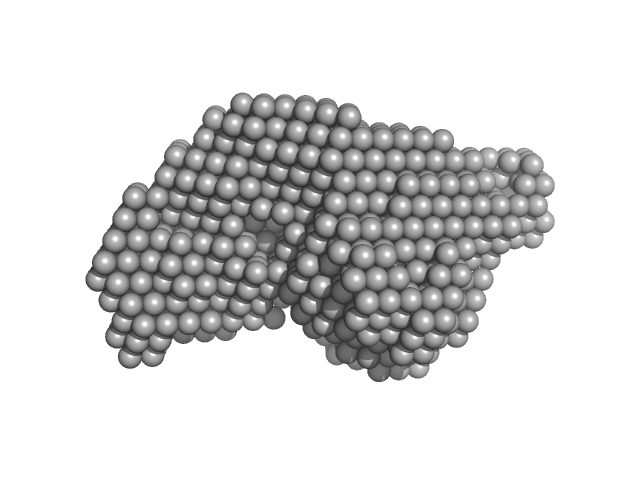
The molecular weight quoted is derived from concentration independent estimates spanning the credibility interval 92–103 kDa. Concentration dependent MW estimates, relative to a BSA standard, is estimated at approximately 82 kDa. Note - the data used to calculate the PDDF, and for model generation, was not normalised to protein concentration (ca. 4.9 mg/ml). The models displayed in this entry include: Top - a single GASBOR model representative and fit; Second from top - an individual refined DAMMIN model that was calculated based on the spatial alignment of a 14 model cohort; Third - a representative SASREF model and; bottom - the fit to the data of an Alphafold2 predicted model that is spatially very similar to the E. coli homologue of the AncB protein (PDB 1L5J; Chain A, RMSD Ca = 0.04 nm). Additional model representatives from GASBOR, DAMMIN and SASREF are made available in the full entry zip archive.
|
|
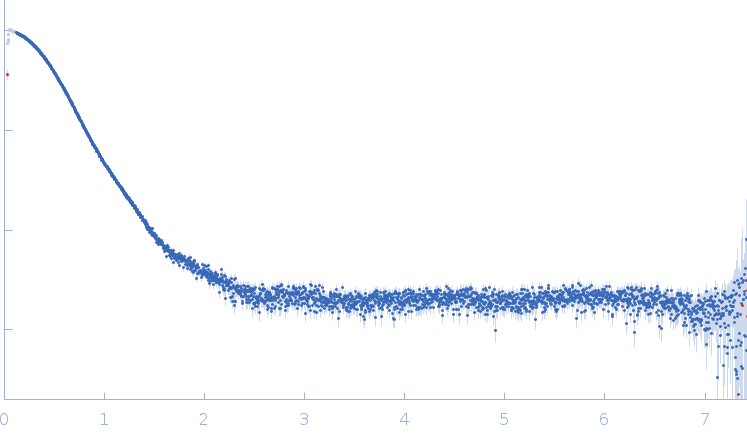 s, nm-1
s, nm-1