|
Synchrotron SAXS data from solutions of calcium-bound calmodulin in 20 mM Hepes, 150 mM NaCl, 2 mM CaCl2 in the presence of a 3-fold excess of CDZ, pH 7.4 were collected on the SWING beam line at the SOLEIL storage ring (Saint-Aubin, France) using a EIGER 4M detector at a sample-detector distance of 2.0 m and at a wavelength of λ = 0.1033 nm (I(s) vs s, where s = 4πsin θ/λ and 2θ is the scattering angle). 1320 successive 1 second frames (0.990 second exposure time / 0.010 second dead time) were collected at 15°C using size-exclusion chromatography SAXS. A 50 μl sample containing 333 µM calmodulin was injected onto a TSKgel G3000SW column (Tosoh Bioscience) and eluted at a 0.50 ml/min flow rate. Scattered intensities were converted into absolute scale (cm-1) values using the scattering of water. Molecular mass was estimated at 16.3kDa using the correlation volume Vc (Rambo, R.P. and Tainer, J. A. (2013) Nature 496(7446):477-81.).
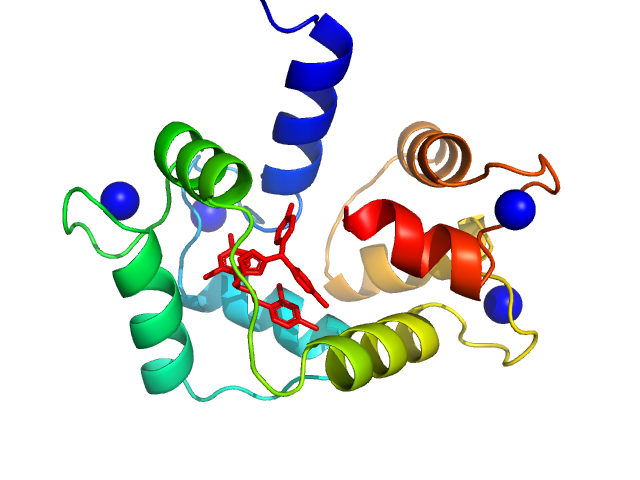
Ab initio model and comparison with the crystal structure:
Top Panel: Comparison of the experimental data (blue dots) with the calculated scattering pattern (red line) of the DENSS density distribution shown on the right. (Grant, T. D. (2018) Nat Methods 15, 191–193).
Bottom panel: Comparison of the experimental data (blue dots) with the calculated scattering pattern (red line) of the crystallographic structure of Calmidazolium-bound Holo-Calmodulin (pdb 7PSZ) obtained using CRYSOL. chi2 is 1.4.
|
|
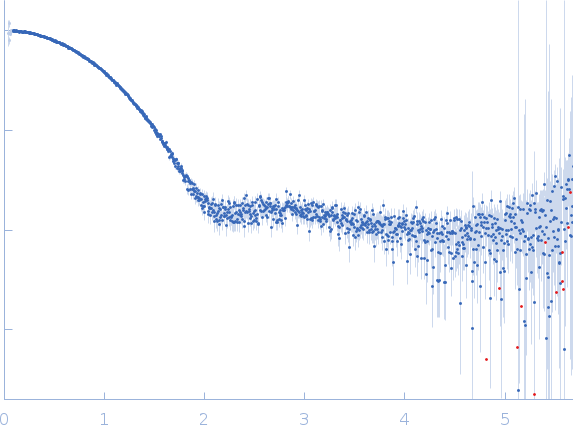 s, nm-1
s, nm-1