|
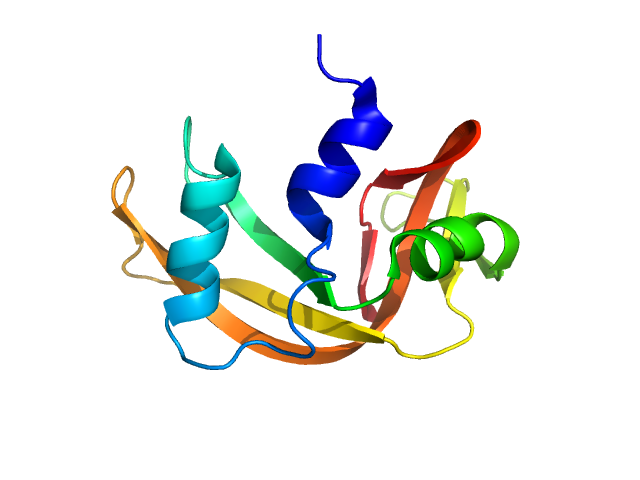
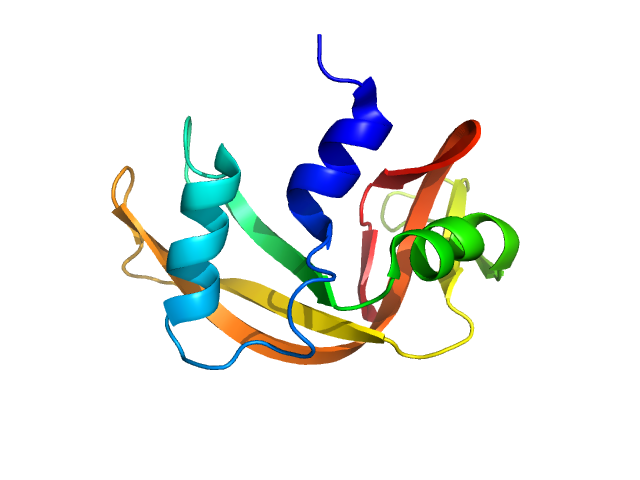
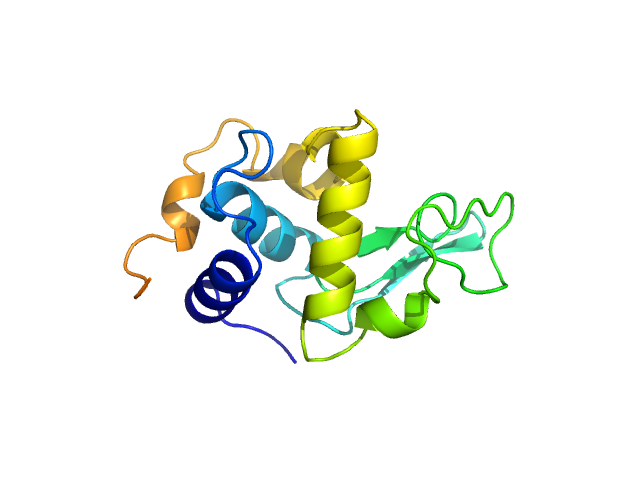
The consensus SANS profile for RNaseA in H2O buffer is a consensus batch+SEC-SANS data set generated using the datcombine tool (ATSAS 3.1.0) with both outlier- and error-filters applied. One SEC-SAXS and five batch data sets were input to datcombine. All contributing data were independent measurements, and no individual measurement was represented more than once in the contributing scattering profile set. The buffer was 50 mM Tris, pH 7.5, 100 mM NaCl. Protein concentrations for batch measurements ranged from 3.6 – 7.7 mg/mL. The ribonuclease A atomistic model for CRYSON and Pepsi-SANS fits was the PDB ID 7RSA with small-molecule crystallisation agents removed. Custom WAXSiS-type calculations (with neutron scattering factors and Gromacs software) used the same coordinates and added explicit waters and appropriate number of ions for the MD calculations.
The data input to datcombine are made available for download in the associated zip file, including also the unsubtracted 6 column format data with resolution data. Model fits are shown in order (top to bottom): CRYSON, Pepsi-SANS, and custom WAXSiS (with neutron scattering factors).
|
|
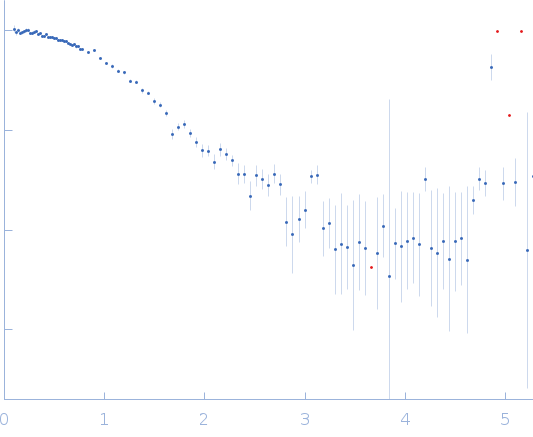 s, nm-1
s, nm-1