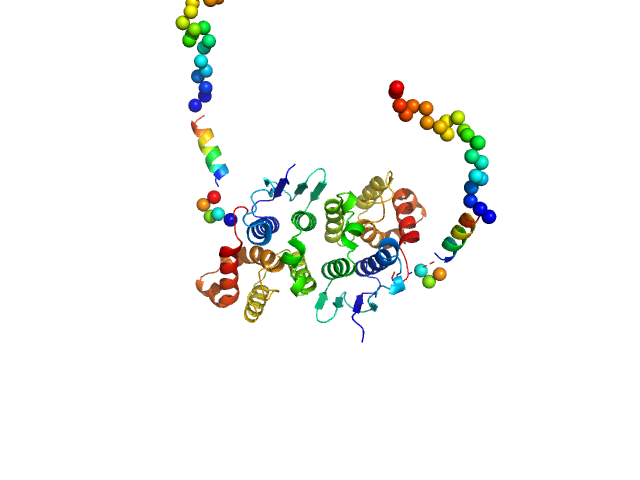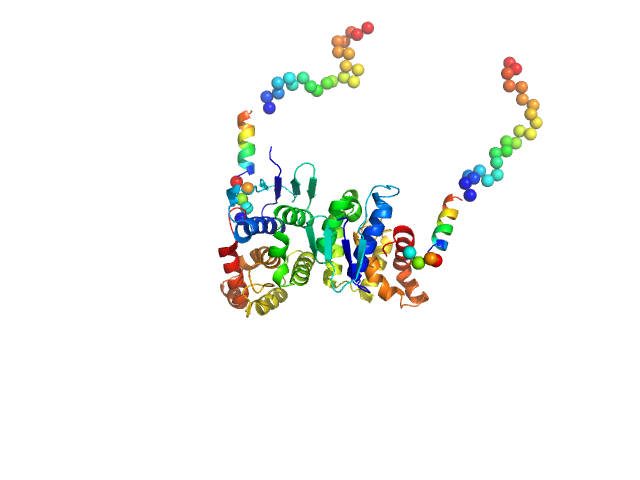|
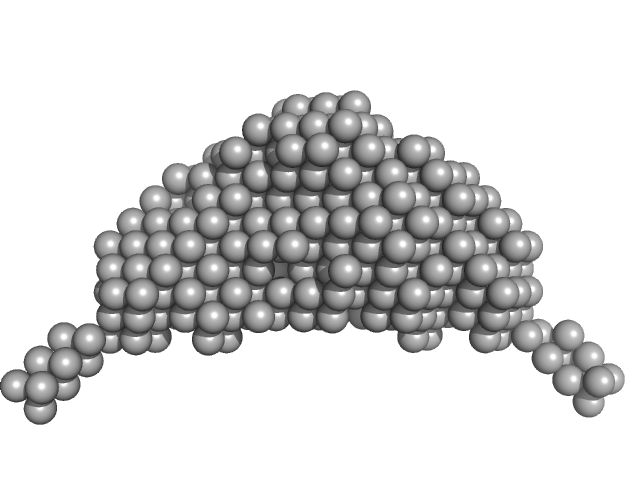
Synchrotron SAXS data of the N-terminal GST-like domain of tRNA import protein (tRip-N) from Plasmodium berghei. The experiment was performed on the SWING beamline at the SOLEIL synchrotron (Saint-Aubin, France) using an EigerX 4M at a sample-detector distance of 2 m and at a wavelength of λ = 0.1033 nm. In-line size-exclusion chromatography was used to separate the sample prior to X-ray exposure. 50 μl of purified recombinant (E. coli) protein at 16 mg/mL were injected on a Bio SEC-3 column (4.6 × 300 mm, 300 Å) equilibrated in 25 mM HEPES-NaOH pH 7.0, 300 mM NaCl, 5% glycerol, 0.005% (m/v) DDM, 5 mM 2-mercaptoethanol at 0.2 mL/min and 15°C. Frames (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle) were collected succesively each 1 s (0.990 s exposure and 0.01 dead time) in two time windows. 180 solvent frames were collected at time 2.5 min prior to the column's void volume and 1230 sample frames were collected from time 9.5 min until the end of the run. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent was subtracted from the sample frames using Foxtrot 3.5.10. Correction for capillary fouling was applied using the US-SOMO 4.0 HPLC SAXS module. Data analysis was performed with BioXTAS RAW 2.1.0. The experimental molecular weight was determined using the method of the volume of correlation. A correction factor has been applied to the Porod volume. MODELLER 10.1 was used to built an atomic model of P. berghei tRip-N based on the crystal structure of P. vivax tRip-N (PDB 5ZKE). CORAL was used to model residues absent in the crystal structure and to test two types of dimeric assemblies of P. berghei tRip-N against SAXS data. The first correspond to a canonical GST-like dimer (above); the second is based on the unusual GST-like dimerization observed in the crystal of P. vivax tRip-N (below). The unusual dimerization provided a slightly better fit to the experimental data.
|
|
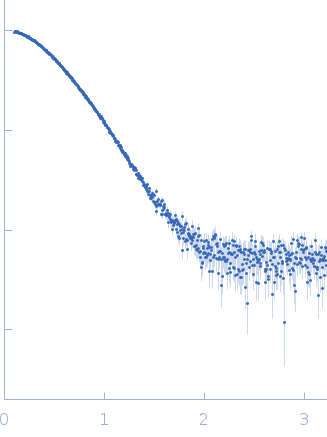 s, nm-1
s, nm-1