|
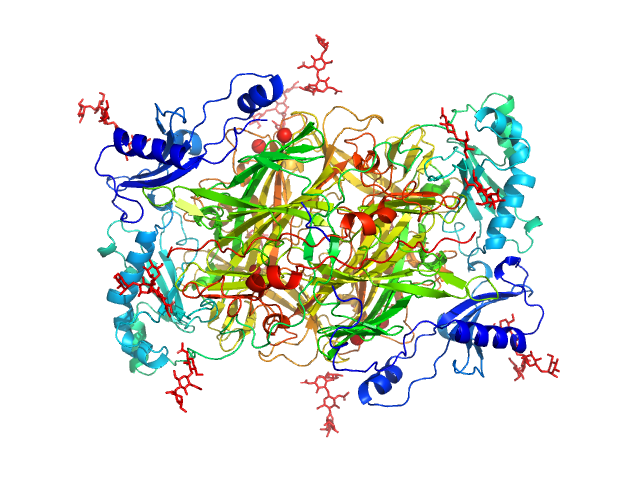
Synchrotron SAXS data from solutions of human AOC3 dimer in 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7.4 were collected on the B21 beam line at the Diamond Light Source (Didcot, UK) using a Pilatus 2M detector at a sample-detector distance of 4 m and at a wavelength of λ = 0.1 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 0.3 and 0.5 mg/ml were measured at 25°C. 27 successive 10 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted and analysed using Primus to produce the data displayed in this entry.
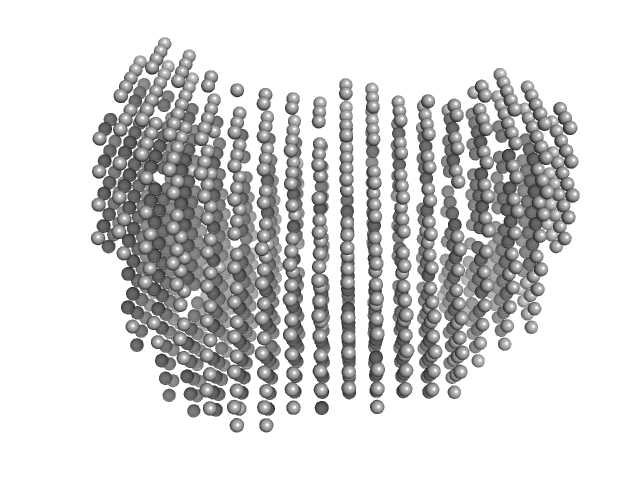
The entry shows 1) the individual ab initio reconstruction and 2) fitting of the experimental scattering curve to the AOC3 crystal structure (PDB entry 8S1Z).
|
|
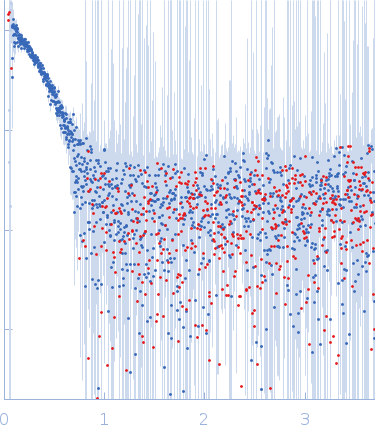 s, nm-1
s, nm-1