| MWexperimental | 49 | kDa |
| MWexpected | 69 | kDa |
| VPorod | 50 | nm3 |
|
log I(s)
3.42×100
3.42×10-1
3.42×10-2
3.42×10-3
|
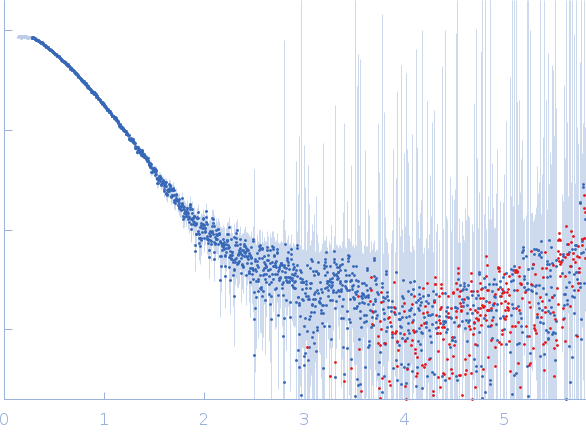 s, nm-1
s, nm-1
|
|
|
|
|
|
|
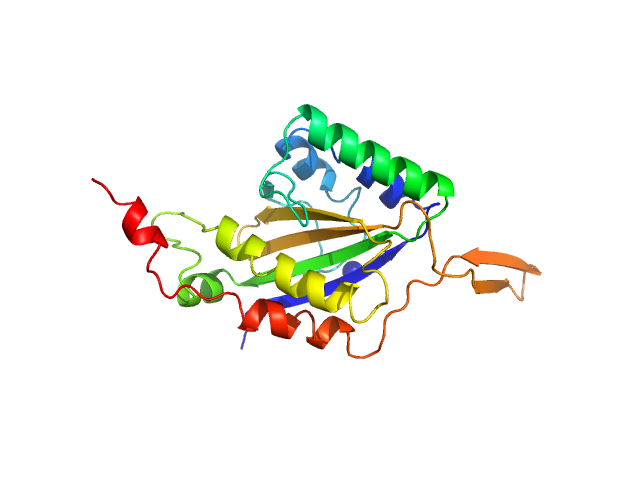
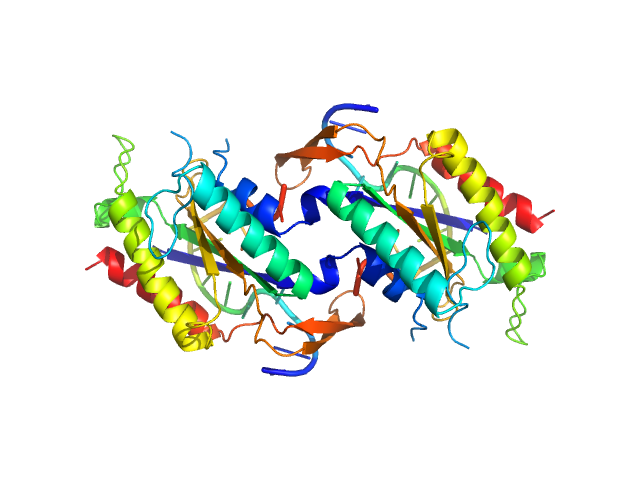
| Synchrotron SAXS data from solutions of the N-terminal domain of relaxase in complex with nic sequence DNA in 20 mM Tris, 300 mM NaCl, 1 mM DTT, pH 8 were collected on the BL11-NCD beam line at ALBA (Cerdanyola del Vallès, Barcelona, Spain) using a ADSC Quantum 210r detector at a sample-detector distance of 2.6 m and at a wavelength of λ = 0.1 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). One solute concentration of 5.00 mg/ml was measured at 25°C. 20 successive 0.100 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. |
|
||||||||||||||||||||||||||||||||||||||||||||||||