|
SAXS data from solutions of farnesoid X receptor homo-dimer bound to inverted repeat DNA in 50 mM HEPES, 150 mM NaCl, 5 % glycerol, 1 mM DTT, 20 µM CDCA, pH 7.4 were collected on a Rigaku BioSAXS-2000 instrument (Pennsylvania State University, University Park, PA, USA) using a Rigaku HyPix-3000 detector at a sample-detector distance of 5.0 m and at a wavelength of λ = 0.154 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). One solute concentration of 0.60 mg/ml was measured at 4°C. Six successive 600 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
Chenodeoxycholic acid (CDCA): https://www.ebi.ac.uk/chembl/explore/compound/CHEMBL240597
|
|
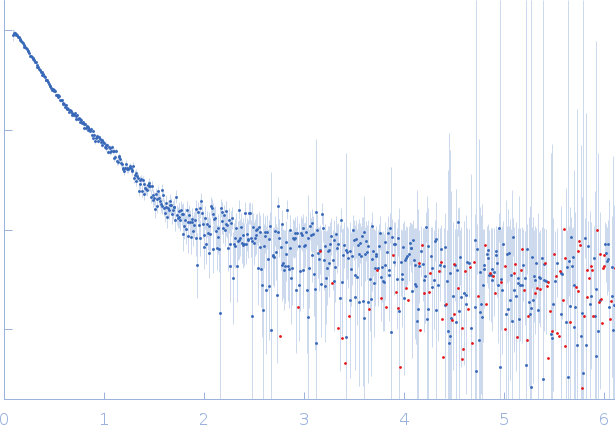 s, nm-1
s, nm-1