Structural basis underlying the synergism of NADase and SLO during group A Streptococcus infection.
Tsai WJ,
Lai YH,
Shi YA,
Hammel M,
Duff AP,
Whitten AE,
Wilde KL,
Wu CM,
Knott R,
Jeng US,
Kang CY,
Hsu CY,
Wu JL,
Tsai PJ,
Chiang-Ni C,
Wu JJ,
Lin YS,
Liu CC,
Senda T,
Wang S
Commun Biol
6(1):124
(2023 Jan 31)
|
|
|
|
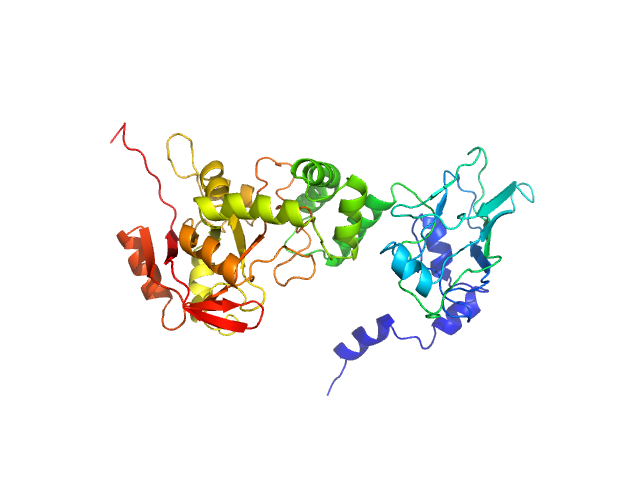
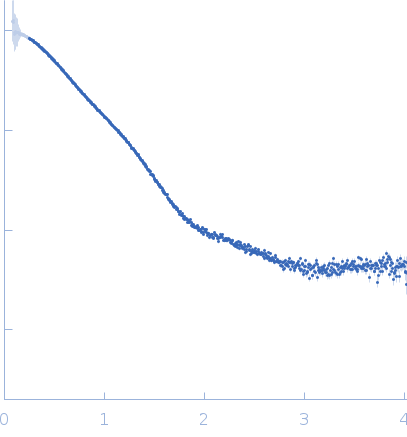
|
| Sample: |
NAD glycohydrolase monomer, 47 kDa Streptococcus pyogenes M1 … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 16
|
|
| RgGuinier |
3.0 |
nm |
| Dmax |
103.0 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
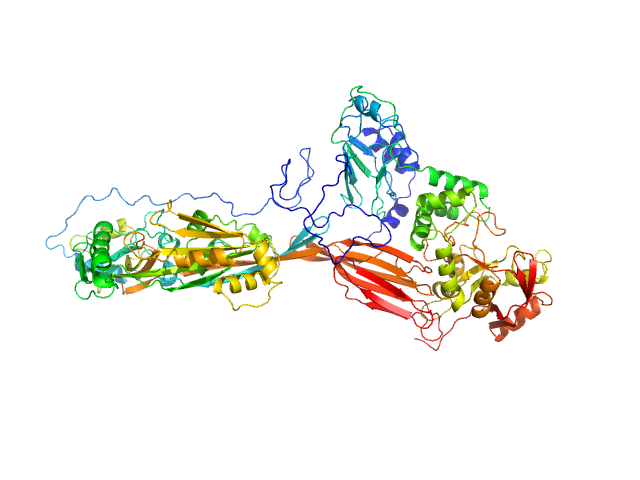
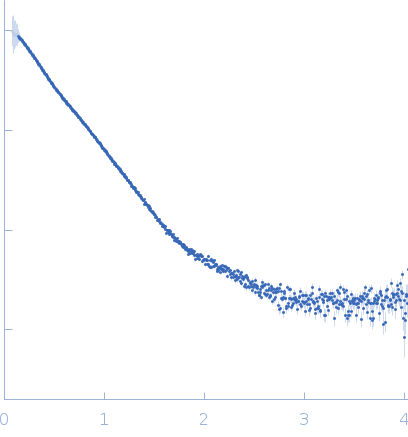
|
| Sample: |
NAD glycohydrolase monomer, 47 kDa Streptococcus pyogenes M1 … protein
Streptolysin O (T66M) monomer, 63 kDa Streptococcus pyogenes serotype … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 16
|
|
| RgGuinier |
4.8 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
125 |
nm3 |
|
|