The Crystal and Solution Structures of Glyceraldehyde-3-phosphate Dehydrogenase Reveal Different Quaternary Structures
Ferreira-da-Silva F,
Pereira P,
Gales L,
Roessle M,
Svergun D,
Moradas-Ferreira P,
Damas A
Journal of Biological Chemistry
281(44):33433-33440
(2006 Nov)
|
|
|
|
|
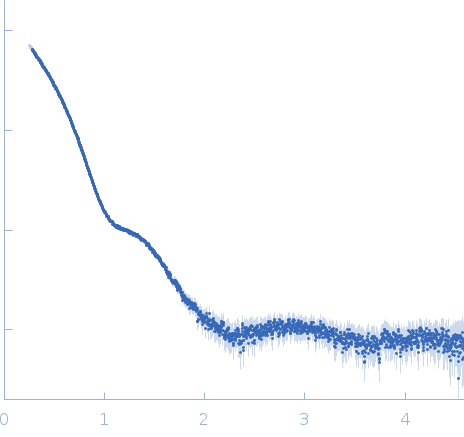
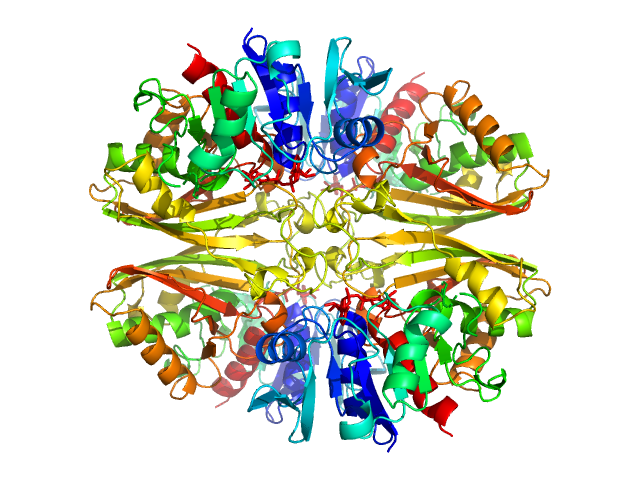
| Sample: |
Glyceraldehyde-3-phosphate dehydrogenase 1 tetramer, 142 kDa Kluyveromyces marxianus protein
|
| Buffer: |
150 mM NaCl, 1 mM beta-mercaptoethanol, 1 mM EDTA, 10 mM TrisHCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 Mar 27
|
|
| RgGuinier |
4.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
234 |
nm3 |
|
|
|
|
|
|
|
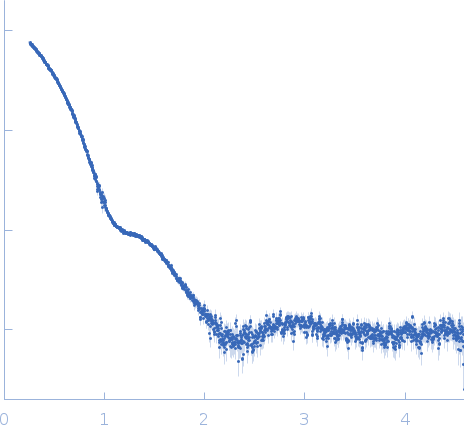
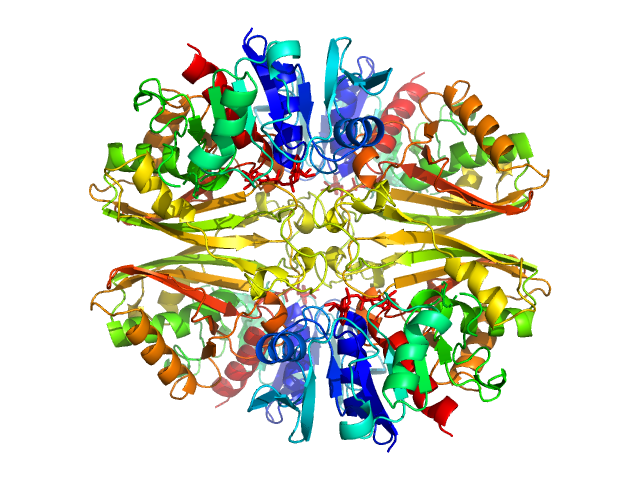
| Sample: |
Glyceraldehyde-3-phosphate dehydrogenase 1 bound to NAD+ tetramer, 142 kDa Kluyveromyces marxianus protein
|
| Buffer: |
150 mM NaCl, 1 mM beta-mercaptoethanol, 1 mM EDTA, 10 mM TrisHCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 Mar 27
|
|
| RgGuinier |
3.7 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
202 |
nm3 |
|
|