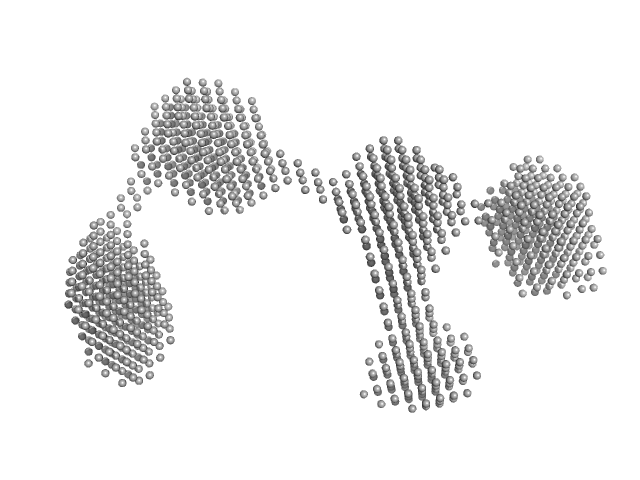Cleavage region organizes the structural architecture of the SINE-derived B2 repressive ribozyme.
Singhal A,
Mrozowich T,
Rivera C,
Chaudhury SN,
Xu L,
Aguilar R,
Badmalia M,
Lee JT,
Patel TR,
Sanbonmatsu KY
Commun Biol
(2026 Mar 23)
|
|
|
|
|
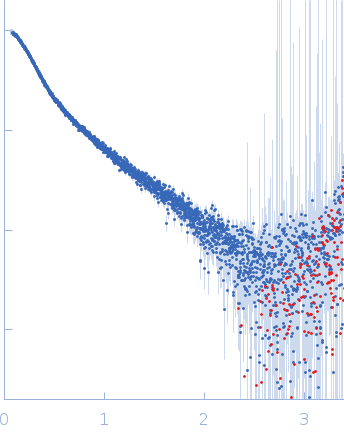
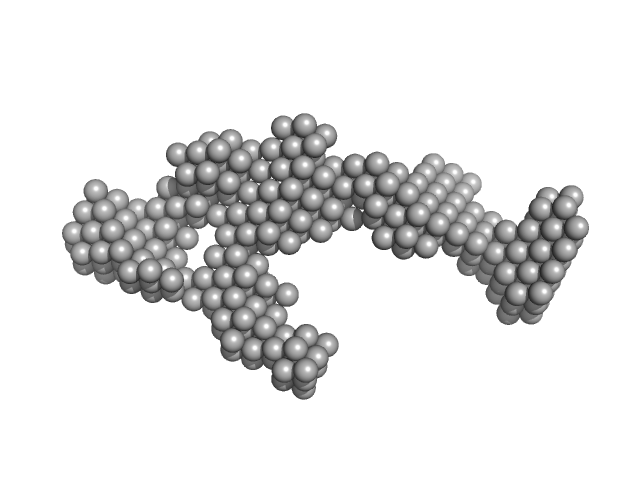
| Sample: |
B2 short interspaced nuclear element (SINE) RNA monomer, 57 kDa Mus musculus RNA
|
| Buffer: |
5 mM Tris-HCl, 10 mM NaCl, 0.01% NP-40, 0.02 mM EDTA, 0.2 mM DTT, 0.5 mM MgCl2 and 1.5 % Glycerol, pH: 7.9 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Feb 20
|
|
| RgGuinier |
5.4 |
nm |
| Dmax |
17.3 |
nm |
| VolumePorod |
131 |
nm3 |
|
|
|
|
|
|
|
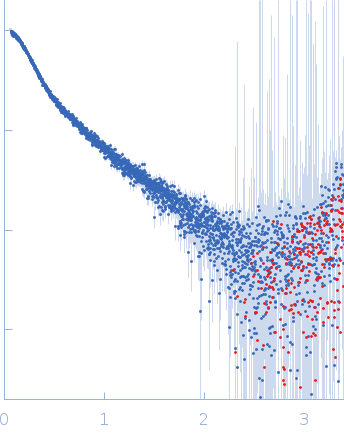
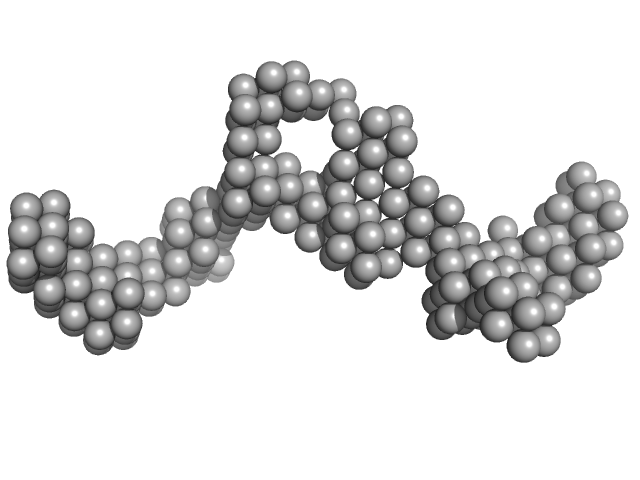
| Sample: |
B2 short interspaced nuclear element (SINE) RNA with two point mutations monomer, 57 kDa Mus musculus RNA
|
| Buffer: |
5 mM Tris-HCl, 10 mM NaCl, 0.01% NP-40, 0.02 mM EDTA, 0.2 mM DTT, 0.5 mM MgCl2 and 1.5 % Glycerol, pH: 7.9 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Feb 20
|
|
| RgGuinier |
5.5 |
nm |
| Dmax |
17.3 |
nm |
| VolumePorod |
136 |
nm3 |
|
|
|
|
|
|

|
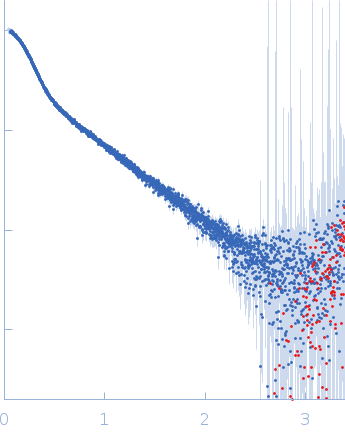
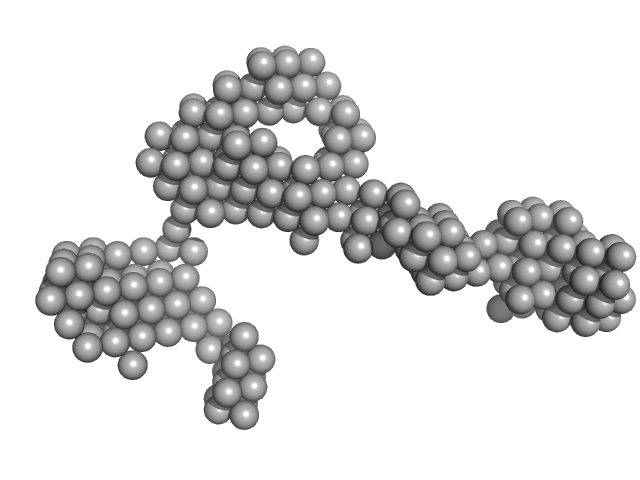
| Sample: |
B2 short interspaced nuclear element (SINE) RNA with deletion mutation 96-105 monomer, 54 kDa Mus musculus RNA
|
| Buffer: |
5 mM Tris-HCl, 10 mM NaCl, 0.01% NP-40, 0.02 mM EDTA, 0.2 mM DTT, 0.5 mM MgCl2 and 1.5 % Glycerol, pH: 7.9 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Dec 18
|
|
| RgGuinier |
5.5 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
135 |
nm3 |
|
|
|
|
|
|
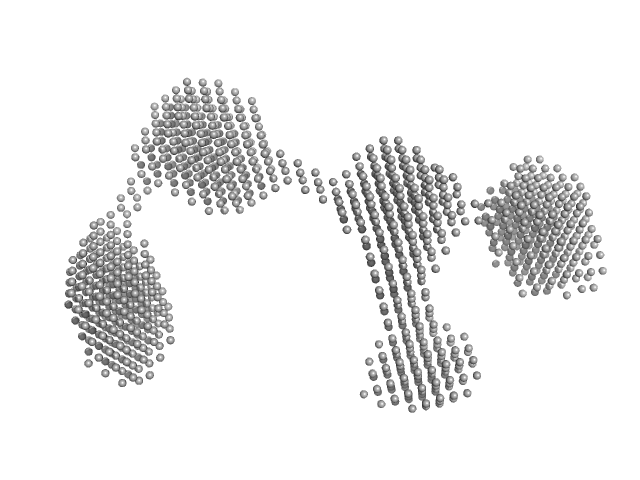
|
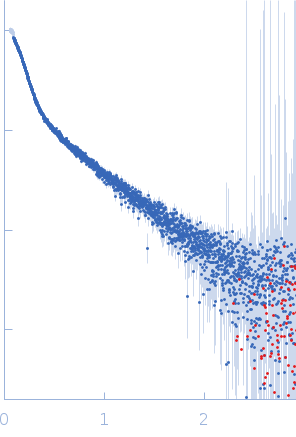
| Sample: |
B2 short interspaced nuclear element (SINE) RNA with deletion mutation 81-124 dimer, 86 kDa Mus musculus RNA
|
| Buffer: |
5 mM Tris-HCl, 10 mM NaCl, 0.01% NP-40, 0.02 mM EDTA, 0.2 mM DTT, 0.5 mM MgCl2 and 1.5 % Glycerol, pH: 7.9 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Aug 19
|
|
| RgGuinier |
8.1 |
nm |
| Dmax |
22.1 |
nm |
|
|