Probing the statistics of sequence-dependent DNA conformations in solution using SAXS.
Koning HJ,
Pullakhandam A,
Whitten AE,
Bond CS,
Peyrard M
Acta Crystallogr D Struct Biol
82(Pt 2):79-99
(2026 Feb 1)
|
|
|
|
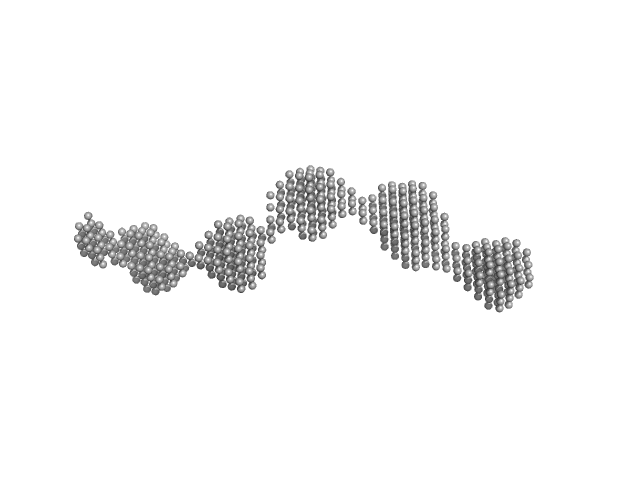

|
| Sample: |
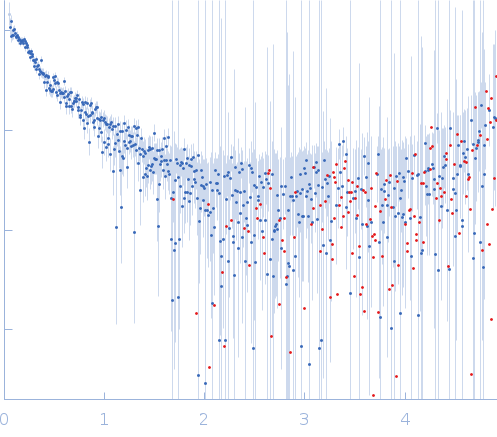
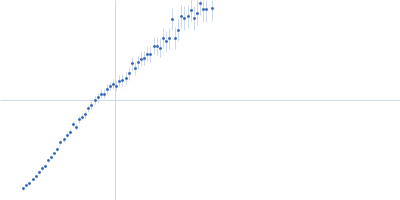
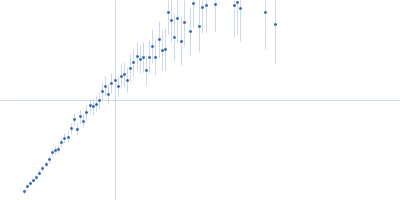
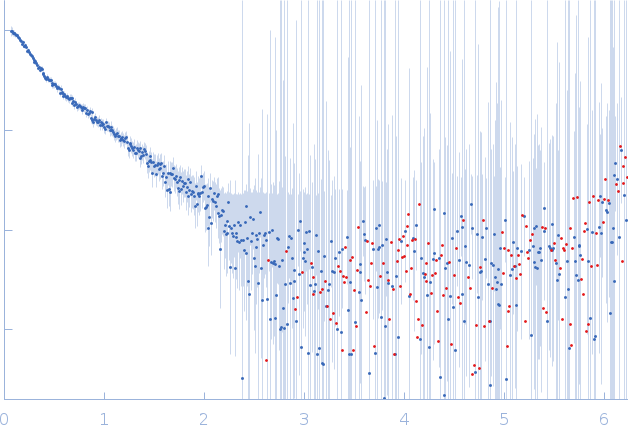
GAGE6 60bp dsDNA oligo monomer, 37 kDa Homo sapiens DNA
|
| Buffer: |
150 mM KCl, 20 mM HEPES, 5% glycerol, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Mar 14
|
|
| RgGuinier |
4.8 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
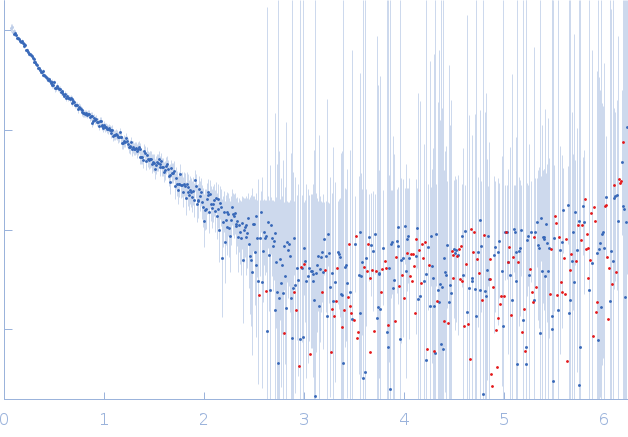
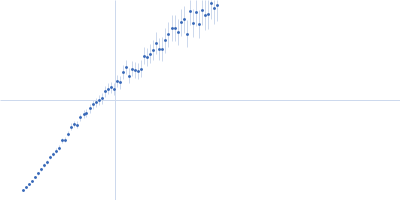
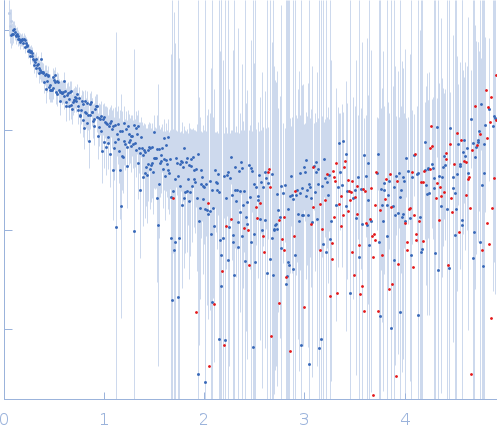
GAGE6_1 dimer, 37 kDa Homo sapiens DNA
|
| Buffer: |
150 mM KCl, 20 mM HEPES, 5% glycerol, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Nov 28
|
|
| RgGuinier |
5.1 |
nm |
| Dmax |
19.2 |
nm |
| VolumePorod |
67 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
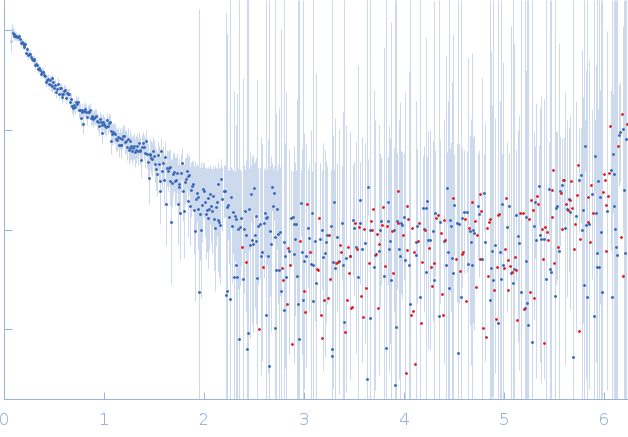
GAGE6_2 dimer, 37 kDa Homo sapiens DNA
|
| Buffer: |
150 mM KCl, 20 mM HEPES, 5% glycerol, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Nov 28
|
|
| RgGuinier |
5.2 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
GAGE6_3 dimer, 37 kDa Homo sapiens DNA
|
| Buffer: |
150 mM KCl, 20 mM HEPES, 5% glycerol, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Nov 28
|
|
| RgGuinier |
5.0 |
nm |
| Dmax |
21.4 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
|
|
|
|
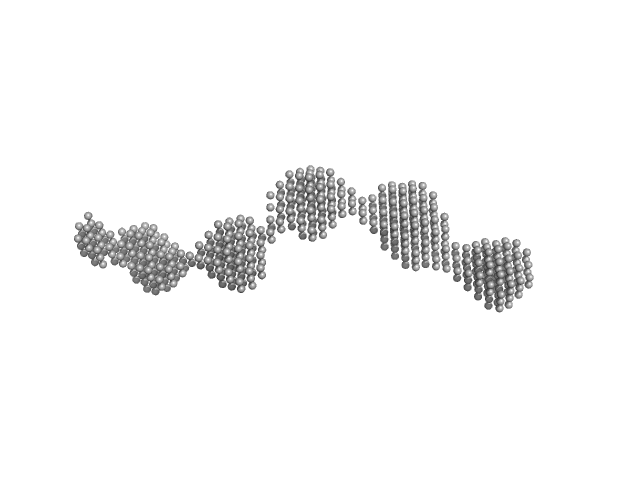

|
| Sample: |
GAGE6 60bp dsDNA oligo monomer, 37 kDa Homo sapiens DNA
|
| Buffer: |
150 mM KCl, 20 mM HEPES, 5% glycerol, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Mar 14
|
|
| RgGuinier |
4.7 |
nm |
| Dmax |
20.7 |
nm |
| VolumePorod |
101 |
nm3 |
|
|