|
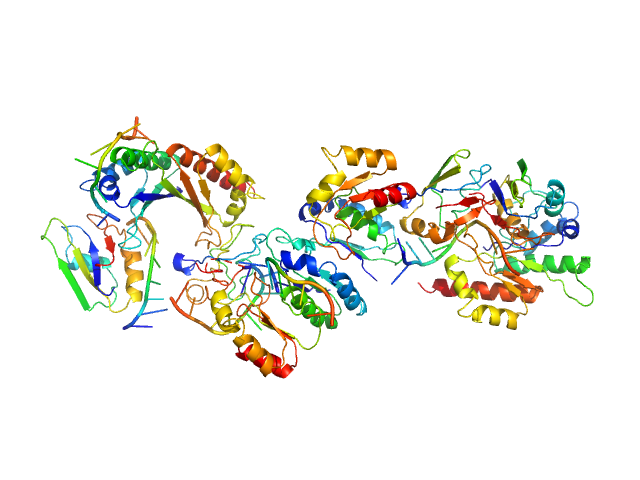
Synchrotron SAXS data from solutions of insulin-like growth factor 2 mRNA-binding protein 2 (IMP2) with RNA in 20 mM HEPES, 2 mM MgCl2, 1 M NaCl, 10% glycerol v/v, 2 mM β-mercaptoethanol, pH 7.4 were collected on the 16-ID (LiX) beam line at the National Synchrotron Light Source II (NSLS-II; Brookhaven National Laboratory, Upton, NY, USA) using dual Pilatus3 S 1M/Pilatus3 900 K detectors at a wavelength of λ = 0.11 nm and 1.5 mm path-length (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed and used to separate proteinaceous species. Samples were loaded into a 96-well auto-sampler plate prior to injection into a pre-equilibrated size exclusion-coupled small-angle X-ray scattering (SEC-SAXS) system. An Agilent 1260 Infinity Bio-Inert high-performance liquid chromatography (HPLC) with an auto-sampler was used for sample injection and SEC. The SEC parameters were as follows: A 100.00 μl sample at 5 mg/ml was injected at a 0.50 ml/min flow rate onto a Cytiva Superdex 200 5/150 column at 4°C. Successive 2 second data frames were collected during the course of 15-minute SEC-elution (15 frames were selected across the sample elution peak). SAXS/WAXS images were radially integrated before being background subtracted using LIX specific python packages py4xs and lixtools. Subtracted profiles were exported and imported for analysis in BioXTAS RAW (RAW).
IMP2 was co-incubated with Cox7b RNA before running the SEC-SAXS experiment. CAUTION: The estimated Porod volume (55.8 nm^3) appears underestimated expected for particles with an experimental MW of 135 kDa. The model of the complex displayed here shows significant steric clashes and discontinuous RNA.
|
|
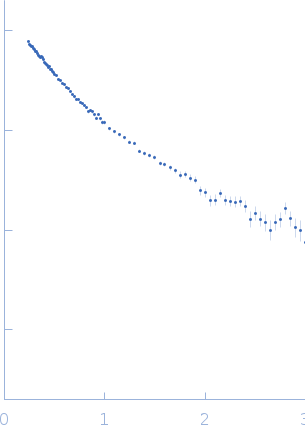 s, nm-1
s, nm-1