|
|
|
|
|
| Sample: |
Hepatitis B virus (HBV) None,
|
| Buffer: |
0.5 M LiCl, 50 mM HEPES, 2 mM DTT, pH 7.5, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Jun 23
|
Coat Protein-Dependent Behavior of Poly(ethylene glycol) Tails in Iron Oxide Core Virus-like Nanoparticles.
ACS Appl Mater Interfaces 7(22):12089-98 (2015)
Malyutin AG, Cheng H, Sanchez-Felix OR, Carlson K, Stein BD, Konarev PV, Svergun DI, Dragnea B, Bronstein LM
|
|
|
|
|
|
|
|
| Sample: |
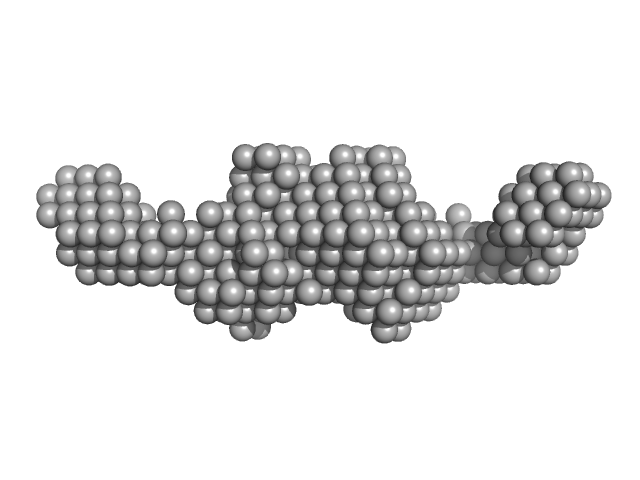
RAID3 dimer, 23 kDa RNA
|
| Buffer: |
PBS 137 mM NaCl 2,7 mM KCl 6,5 mM Na2HPO4 1,5 mM KH2PO4, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2013 Oct 31
|
RAID3--An interleukin-6 receptor-binding aptamer with post-selective modification-resistant affinity.
RNA Biol 12(9):1043-53 (2015)
Mittelberger F, Meyer C, Waetzig GH, Zacharias M, Valentini E, Svergun DI, Berg K, Lorenzen I, Grötzinger J, Rose-John S, Hahn U
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
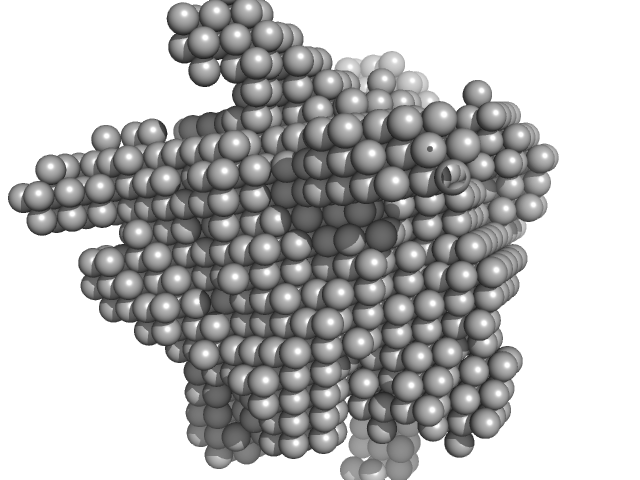
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - 5α-cholestan-3-one) without Dox monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
6.1 |
nm |
| Dmax |
20.5 |
nm |
|
|
|
|
|
|

|
| Sample: |
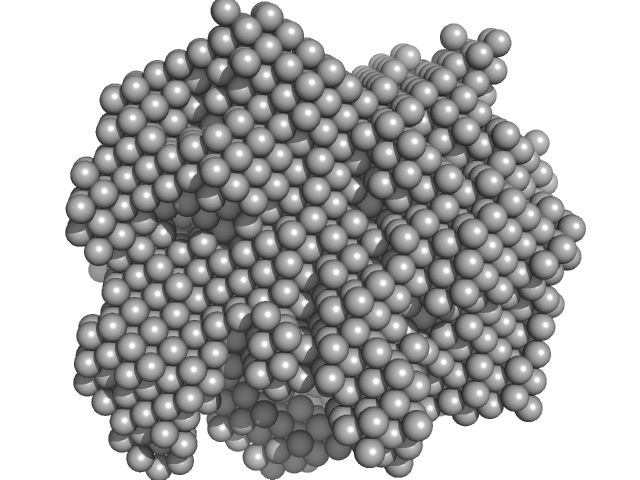
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - 5α-cholestan-3-one) with Dox (10%) monomer, 220 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
7.5 |
nm |
| Dmax |
25.5 |
nm |
|
|
|
|
|
|
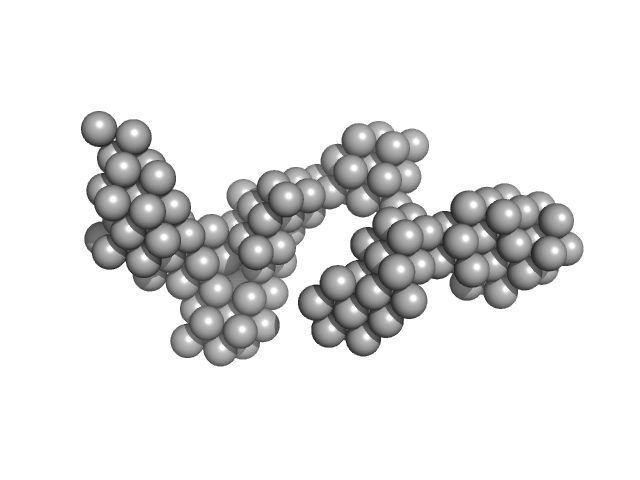
|
| Sample: |
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - Opb-Chol) without Dox monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
6.4 |
nm |
| Dmax |
22.5 |
nm |
|
|
|
|
|
|
|
| Sample: |
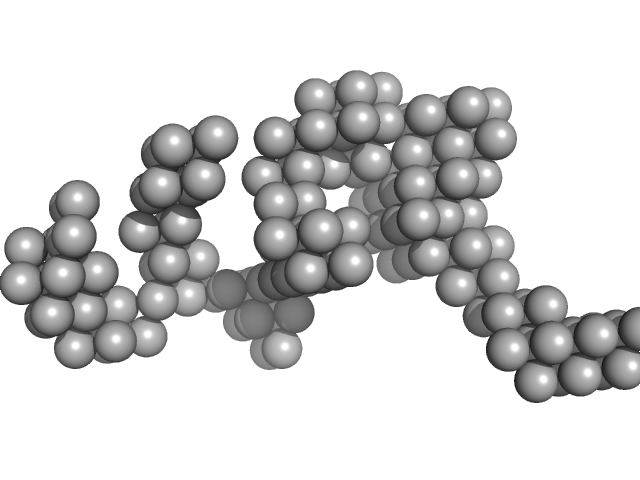
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - Opb-Chol) with Dox (10%) monomer, 225 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
10.0 |
nm |
| Dmax |
21.9 |
nm |
|
|
|
|
|
|
|
| Sample: |
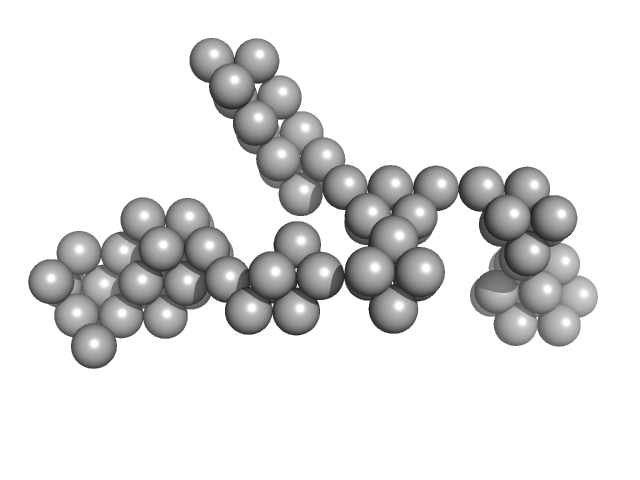
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - Lev-Chol) without Dox monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
6.5 |
nm |
| Dmax |
22.9 |
nm |
|
|
|
|
|
|
|
| Sample: |
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - Lev-Chol) with Dox (10%) monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
6.9 |
nm |
| Dmax |
23.8 |
nm |
|
|
|
|
|
|
|
| Sample: |
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - cholest-4-en-3-one) without Dox monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 16
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
5.9 |
nm |
| Dmax |
19.7 |
nm |
|
|
|
|
|
|
|
| Sample: |
Hydrolytically Degradable Polymer Micelles for Drug Delivery (structure of hydrophobic substituent - cholest-4-en-3-one) with Dox (10%) monomer, 125 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Aug 19
|
Hydrolytically degradable polymer micelles for drug delivery: a SAXS/SANS kinetic study.
Biomacromolecules 14(11):4061-70 (2013)
Filippov SK, Franklin JM, Konarev PV, Chytil P, Etrych T, Bogomolova A, Dyakonova M, Papadakis CM, Radulescu A, Ulbrich K, Stepanek P, Svergun DI
|
| RgGuinier |
5.5 |
nm |
| Dmax |
18.3 |
nm |
|
|