|
|
|
|

|
| Sample: |
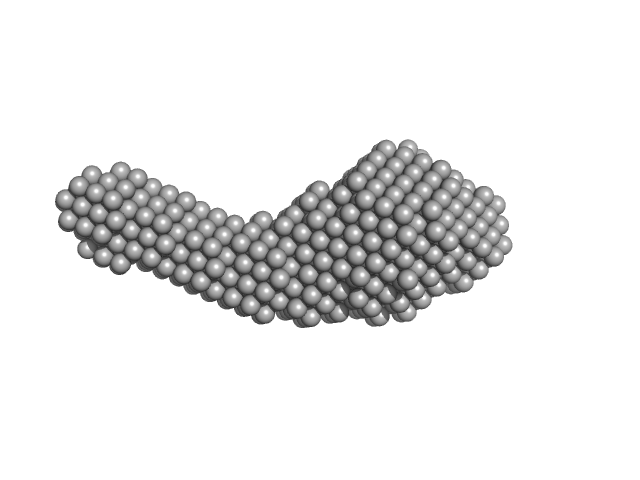
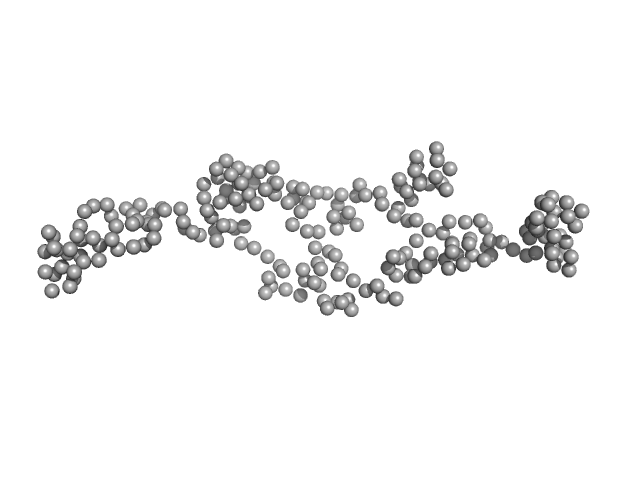
Guanylate-binding protein 1 monomer, 68 kDa Homo sapiens protein
|
| Buffer: |
50 mM TRIS, 5 mM MgCl2, 150 mM NaCl, pH: 7.9
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Apr 30
|
Integrative dynamic structural biology unveils conformers essential for the oligomerization of a large GTPase.
Elife 12 (2023)
Peulen TO, Hengstenberg CS, Biehl R, Dimura M, Lorenz C, Valeri A, Folz J, Hanke CA, Ince S, Vöpel T, Farago B, Gohlke H, Klare JP, Stadler AM, Seidel CAM, Herrmann C
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.4 |
nm |
| VolumePorod |
105 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
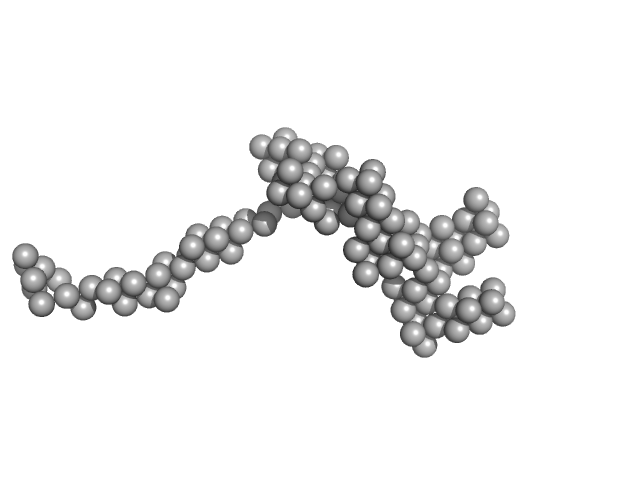
AT5g04600/T32M21_200 monomer, 25 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HNa2PO4, 300 mM NaCl, 5% glycerol (v/v), 1 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 22
|
Structural and functional analysis of a plant nucleolar RNA chaperone-like protein.
Sci Rep 13(1):9656 (2023)
Fernandes R, Ostendorp A, Ostendorp S, Mehrmann J, Falke S, Graewert MA, Weingartner M, Kehr J, Hoth S
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Receptor-type tyrosine-protein phosphatase alpha dimer, 150 kDa Homo sapiens protein
|
| Buffer: |
10 mM phosphate, 137 mM NaCl, 2.7 mM KCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2021 Aug 19
|
High Density of N- and O-Glycosylation Shields and Defines the Structural Dynamics of the Intrinsically Disordered Ectodomain of Receptor-type Protein Tyrosine Phosphatase Alpha
JACS Au (2023)
Chien Y, Wang Y, Sridharan D, Kuo C, Chien C, Uchihashi T, Kato K, Angata T, Meng T, Hsu S, Khoo K
|
| RgGuinier |
8.3 |
nm |
| Dmax |
35.3 |
nm |
|
|
|
|
|
|

|
| Sample: |
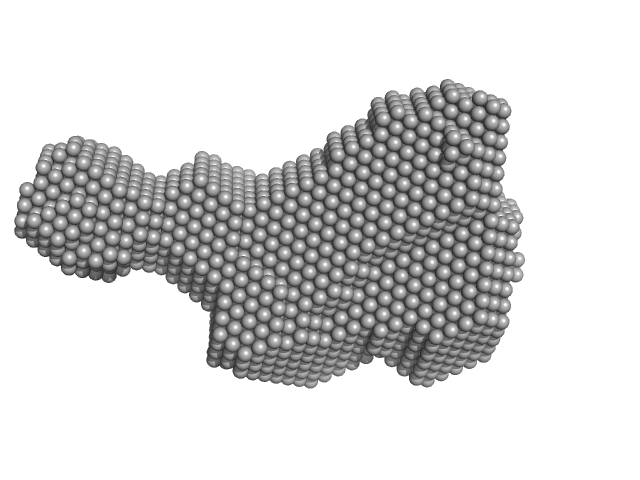
3-phosphoinositide-dependent protein kinase 1 monomer, 65 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl pH 7.4, 250 mM NaCl, 1 mM DTT, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 26
|
Modulation of the substrate specificity of the kinase PDK1 by distinct conformations of the full-length protein
Science Signaling 16(789) (2023)
Sacerdoti M, Gross L, Riley A, Zehnder K, Ghode A, Klinke S, Anand G, Paris K, Winkel A, Herbrand A, Godage H, Cozier G, Süß E, Schulze J, Pastor-Flores D, Bollini M, Cappellari M, Svergun D, Gräwert ...
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.2 |
nm |
| VolumePorod |
107 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
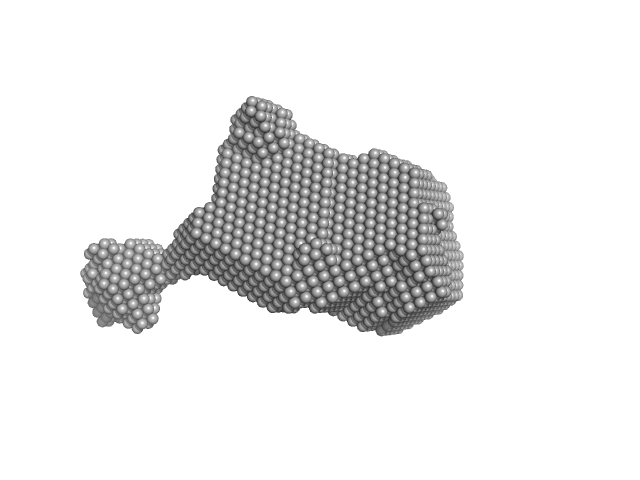
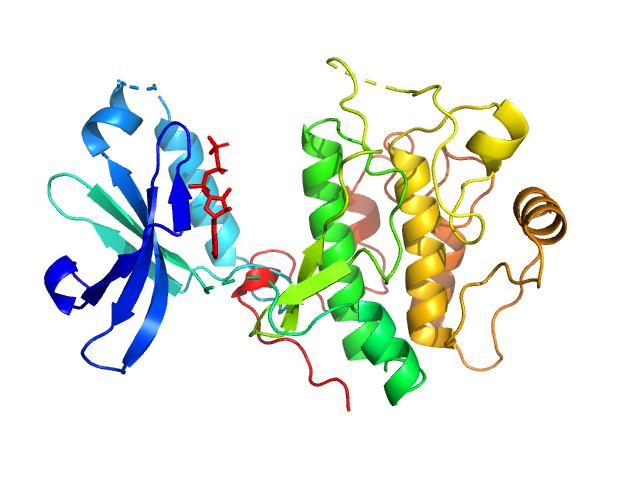
3-phosphoinositide-dependent protein kinase 1 monomer, 65 kDa Homo sapiens protein
2-O-benzoyl-Ins(1,3,4,5,6)P5 monomer, 1 kDa synthetic construct
|
| Buffer: |
20 mM Tris-HCl pH 7.4, 250 mM NaCl, 1 mM DTT, 1 μM HYG8, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 26
|
Modulation of the substrate specificity of the kinase PDK1 by distinct conformations of the full-length protein
Science Signaling 16(789) (2023)
Sacerdoti M, Gross L, Riley A, Zehnder K, Ghode A, Klinke S, Anand G, Paris K, Winkel A, Herbrand A, Godage H, Cozier G, Süß E, Schulze J, Pastor-Flores D, Bollini M, Cappellari M, Svergun D, Gräwert ...
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
98 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
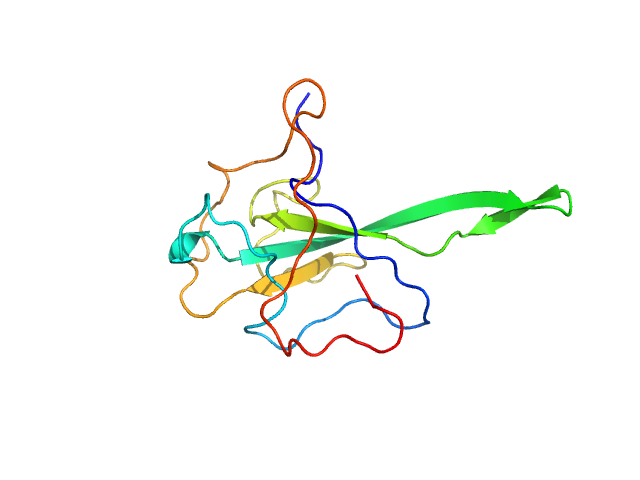
3-phosphoinositide-dependent protein kinase 1 (Y188G Q292A) monomer, 35 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl pH 7.4, 250 mM NaCl, 1 mM DTT, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 26
|
Modulation of the substrate specificity of the kinase PDK1 by distinct conformations of the full-length protein
Science Signaling 16(789) (2023)
Sacerdoti M, Gross L, Riley A, Zehnder K, Ghode A, Klinke S, Anand G, Paris K, Winkel A, Herbrand A, Godage H, Cozier G, Süß E, Schulze J, Pastor-Flores D, Bollini M, Cappellari M, Svergun D, Gräwert ...
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Nucleoprotein monomer, 15 kDa Severe acute respiratory … protein
|
| Buffer: |
25 mM potassium phosphate, 150 mM KCl, 2 mM TCEP, pH: 6.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Aug 16
|
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5'-genomic RNA elements.
Nat Commun 14(1):3331 (2023)
Korn SM, Dhamotharan K, Jeffries CM, Schlundt A
|
| RgGuinier |
1.6 |
nm |
| Dmax |
6.1 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
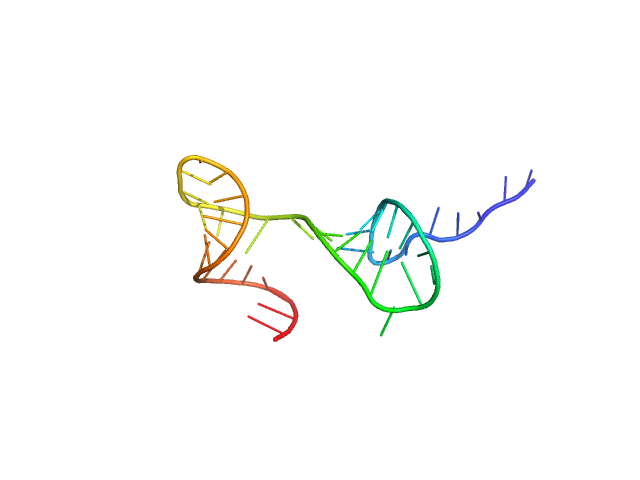
Stem loop 2 and 3 in the 5'-genomic end of SARS-CoV-2 monomer, 14 kDa Severe acute respiratory … RNA
|
| Buffer: |
25 mM potassium phosphate, 150 mM KCl, 2 mM TCEP, pH: 6.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Nov 22
|
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5'-genomic RNA elements.
Nat Commun 14(1):3331 (2023)
Korn SM, Dhamotharan K, Jeffries CM, Schlundt A
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
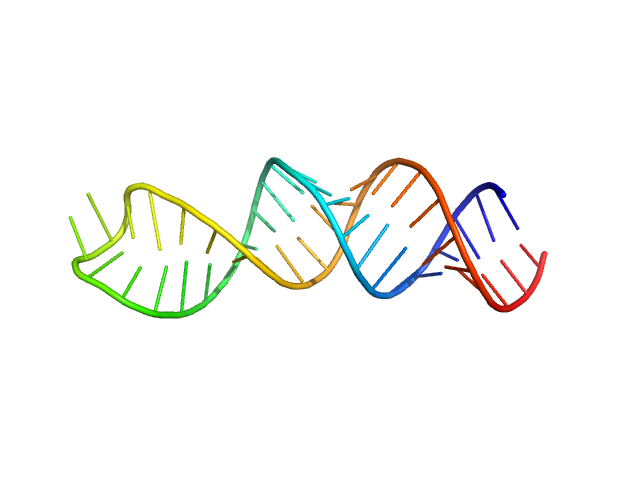
Stem loop 4 in the 5'-genomic end of SARS-CoV-2 monomer, 14 kDa Severe acute respiratory … RNA
|
| Buffer: |
25 mM potassium phosphate, 150 mM KCl, 2 mM TCEP, pH: 6.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Nov 22
|
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5'-genomic RNA elements.
Nat Commun 14(1):3331 (2023)
Korn SM, Dhamotharan K, Jeffries CM, Schlundt A
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
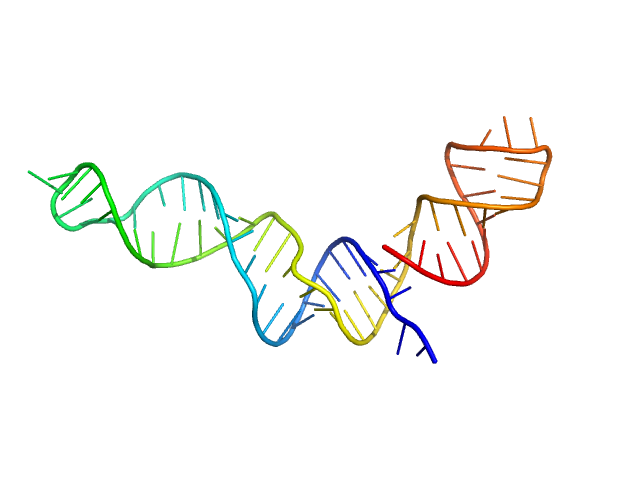
Stem loop 4 with AU extension in the 5'-genomic end of SARS-CoV-2 monomer, 22 kDa Severe acute respiratory … RNA
|
| Buffer: |
25 mM potassium phosphate, 150 mM KCl, 2 mM TCEP, pH: 6.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Nov 22
|
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5'-genomic RNA elements.
Nat Commun 14(1):3331 (2023)
Korn SM, Dhamotharan K, Jeffries CM, Schlundt A
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
33 |
nm3 |
|
|