|
|
|
|
|
| Sample: |
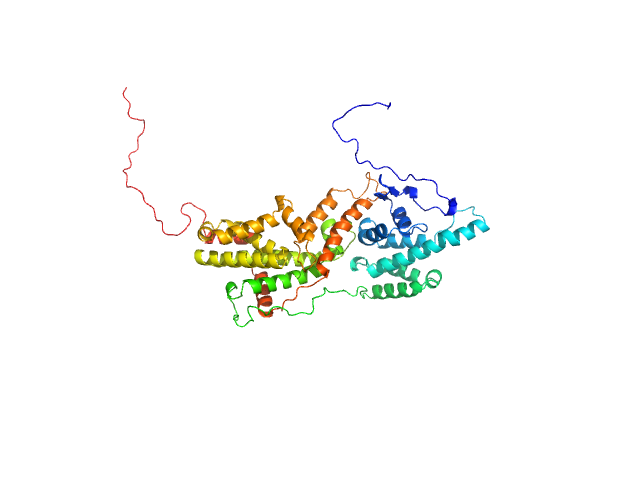
Cell division control protein 25 monomer, 55 kDa Candida albicans (strain … protein
|
| Buffer: |
20 mM Na-phosphate, 150 mM NaCl, 5% glycerol, 3 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 21
|
Pathogen-specific structural features of Candida albicans Ras1 activation complex: uncovering new antifungal drug targets.
mBio :e0063823 (2023)
Manso JA, Carabias A, Sárkány Z, de Pereda JM, Pereira PJB, Macedo-Ribeiro S
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
81 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
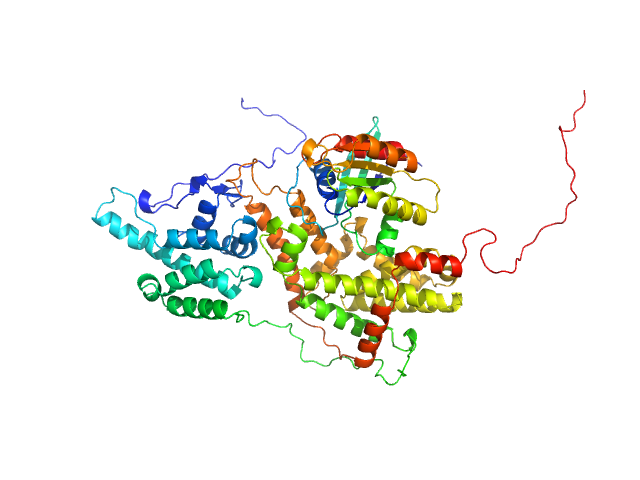
GTP-binding domain of Ras-like protein 1 monomer, 19 kDa protein
Cell division control protein 25 monomer, 55 kDa Candida albicans (strain … protein
|
| Buffer: |
20 mM Na-phosphate, 150 mM NaCl, 1 mM EDTA, 5% glycerol, 3 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 21
|
Pathogen-specific structural features of Candida albicans Ras1 activation complex: uncovering new antifungal drug targets.
mBio :e0063823 (2023)
Manso JA, Carabias A, Sárkány Z, de Pereda JM, Pereira PJB, Macedo-Ribeiro S
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
99 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ras-like protein 1 monomer, 33 kDa Candida albicans protein
Cell division control protein 25 monomer, 55 kDa Candida albicans (strain … protein
|
| Buffer: |
20 mM Na-phosphate, 150 mM NaCl, 1 mM EDTA, 5% glycerol, 3 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 21
|
Pathogen-specific structural features of Candida albicans Ras1 activation complex: uncovering new antifungal drug targets.
mBio :e0063823 (2023)
Manso JA, Carabias A, Sárkány Z, de Pereda JM, Pereira PJB, Macedo-Ribeiro S
|
| RgGuinier |
4.0 |
nm |
| Dmax |
17.7 |
nm |
| VolumePorod |
149 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
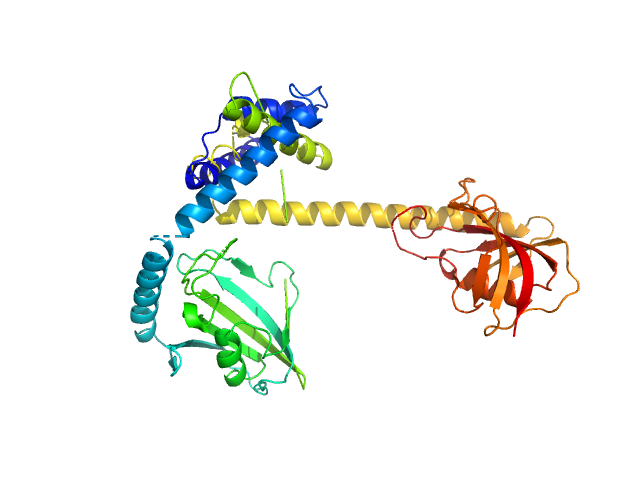
Outer membrane protein MIP dimer, 46 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 10 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 16
|
Legionella pneumophila macrophage infectivity potentiator protein appendage domains modulate protein dynamics and inhibitor binding
International Journal of Biological Macromolecules :126366 (2023)
Wiedemann C, Whittaker J, Carrillo V, Goretzki B, Dajka M, Tebbe F, Harder J, Krajczy P, Joseph B, Hausch F, Guskov A, Hellmich U
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
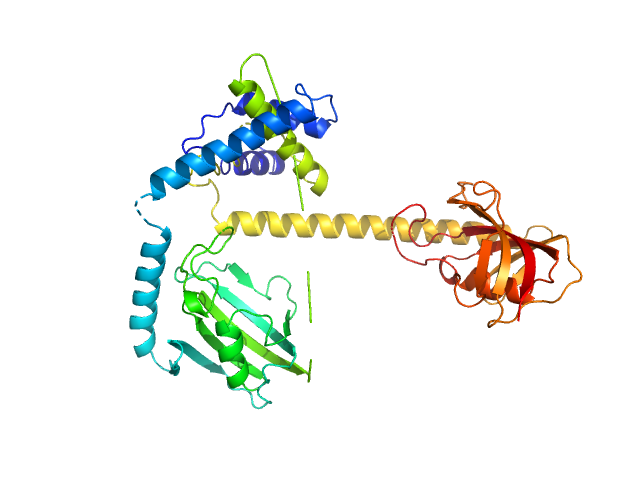
Outer membrane protein MIP dimer, 46 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 10 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 16
|
Legionella pneumophila macrophage infectivity potentiator protein appendage domains modulate protein dynamics and inhibitor binding
International Journal of Biological Macromolecules :126366 (2023)
Wiedemann C, Whittaker J, Carrillo V, Goretzki B, Dajka M, Tebbe F, Harder J, Krajczy P, Joseph B, Hausch F, Guskov A, Hellmich U
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
65 |
nm3 |
|
|
|
|
|
|
|
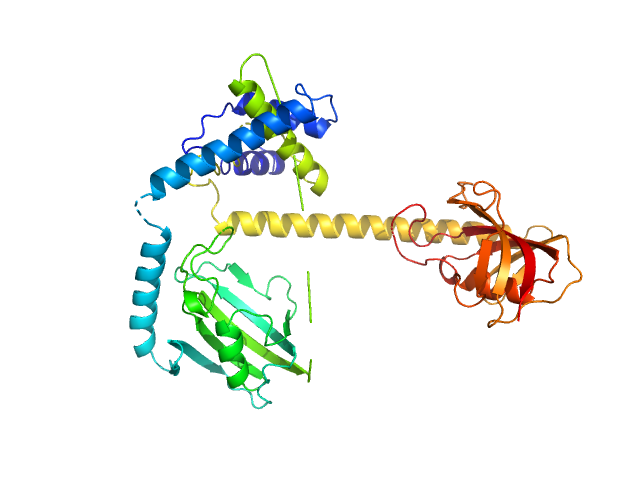
| Sample: |
Outer membrane protein MIP dimer, 46 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 10 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 16
|
Legionella pneumophila macrophage infectivity potentiator protein appendage domains modulate protein dynamics and inhibitor binding
International Journal of Biological Macromolecules :126366 (2023)
Wiedemann C, Whittaker J, Carrillo V, Goretzki B, Dajka M, Tebbe F, Harder J, Krajczy P, Joseph B, Hausch F, Guskov A, Hellmich U
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Xist A-repeat lncRNA monomer, 148 kDa Homo sapiens RNA
|
| Buffer: |
25 mM K-HEPES, 0.1 mM Na-EDTA, 150 mM KCl, 15 mM MgCl2, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 20
|
Conformation and structural dynamics of the Xist lncRNA A-repeats
(2023)
Jones A, Gabel F, Bohn S, Wolfe G, Sattler M
|
| RgGuinier |
12.1 |
nm |
| Dmax |
45.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
Xist A-repeat lncRNA monomer, 148 kDa Homo sapiens RNA
|
| Buffer: |
100 mM HEPES, 100 mM NaCl, 10 mM MgCl2, pH: 8
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 20
|
Conformation and structural dynamics of the Xist lncRNA A-repeats
(2023)
Jones A, Gabel F, Bohn S, Wolfe G, Sattler M
|
| RgGuinier |
9.8 |
nm |
| Dmax |
40.0 |
nm |
| VolumePorod |
657 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Xist A-repeat lncRNA monomer, 148 kDa Homo sapiens RNA
|
| Buffer: |
20 mM HEPES-KOH, 100 mM KCl, 0.2 mM EDTA, 0.5 mM DTT, 0.5 mM PMSF, 20% glycerol, 3.25 mM MgCl2, pH: 7.9
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 20
|
Conformation and structural dynamics of the Xist lncRNA A-repeats
(2023)
Jones A, Gabel F, Bohn S, Wolfe G, Sattler M
|
| RgGuinier |
8.9 |
nm |
| Dmax |
35.0 |
nm |
| VolumePorod |
414 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Xist A-repeat lncRNA monomer, 148 kDa Homo sapiens RNA
|
| Buffer: |
10 mM NaH2PO4/Na2HPO4, 100 mM NaCl, 0.02 mM EDTA, 0.02% azide, pH: 6
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 20
|
Conformation and structural dynamics of the Xist lncRNA A-repeats
(2023)
Jones A, Gabel F, Bohn S, Wolfe G, Sattler M
|
| RgGuinier |
7.6 |
nm |
| Dmax |
30.0 |
nm |
| VolumePorod |
370 |
nm3 |
|
|