|
|
|
|
|
| Sample: |
RNA Short Tetraloop Hairpin Triplex monomer, 17 kDa RNA
|
| Buffer: |
200 mM KCl, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.6 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Short Tetraloop Hairpin Triplex monomer, 17 kDa RNA
|
| Buffer: |
5 mM MgCl2, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.2 |
nm |
| Dmax |
10.2 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Duplex monomer, 21 kDa RNA
|
| Buffer: |
200 mM NaCl, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.8 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Duplex monomer, 21 kDa RNA
|
| Buffer: |
200 mM KCl, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Duplex monomer, 21 kDa RNA
|
| Buffer: |
5 mM MgCl2, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.3 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Triplex monomer, 28 kDa RNA
|
| Buffer: |
200 mM NaCl, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.3 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Triplex monomer, 28 kDa RNA
|
| Buffer: |
200 mM KCl, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.3 |
nm |
|
|
|
|
|
|
|
| Sample: |
RNA Long Tetraloop Hairpin Triplex monomer, 28 kDa RNA
|
| Buffer: |
5 mM MgCl2, 20 mM Na-MES, 50 μM EDTA, pH: 5.5
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2019 Sep 20
|
RNA triplex structures revealed by WAXS-driven MD simulations
(2022)
Chen Y, He W, Kirmizialtin S, Pollack L
|
| RgGuinier |
2.9 |
nm |
| Dmax |
18.2 |
nm |
|
|
|
|
|
|
|
| Sample: |
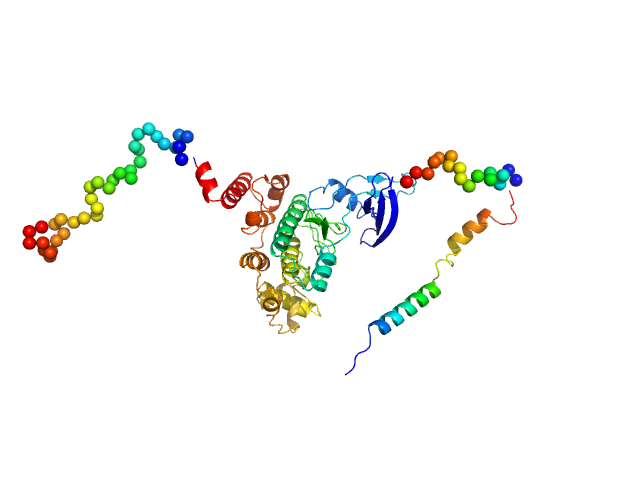
Glycogen synthase kinase 3 monomer, 52 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
20 mM Tris pH 8.0, 100 mM NaCl, 0.5 mM TCEP, pH: 8
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Nov 19
|
N-terminal phosphorylation regulates the activity of glycogen synthase kinase 3 from Plasmodium falciparum.
Biochem J 479(3):337-356 (2022)
Pazicky S, Alder A, Mertens H, Svergun D, Gilberger T, Löw C
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
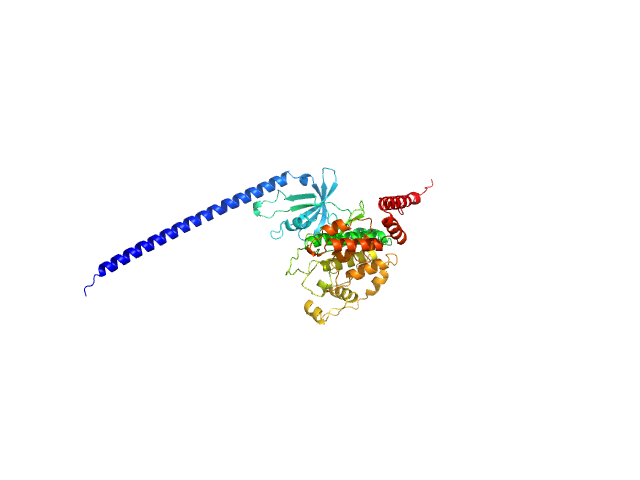
Glycogen synthase kinase 3 monomer, 52 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
20 mM Tris pH 8.0, 100 mM NaCl, 0.5 mM TCEP, pH: 8
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Nov 19
|
N-terminal phosphorylation regulates the activity of glycogen synthase kinase 3 from Plasmodium falciparum.
Biochem J 479(3):337-356 (2022)
Pazicky S, Alder A, Mertens H, Svergun D, Gilberger T, Löw C
|
| RgGuinier |
3.3 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
102 |
nm3 |
|
|