|
|
|
|
|
| Sample: |
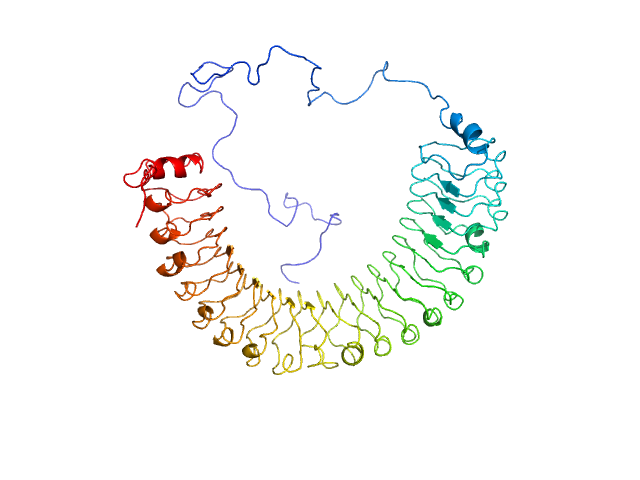
Leucine-rich repeat protein SHOC-2 monomer, 65 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris pH 7.5, 100 mM NaCl, 1 mM TCEP, 1 mM MnCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Jun 20
|
Structural basis for SHOC2 modulation of RAS signalling.
Nature (2022)
Liau NPD, Johnson MC, Izadi S, Gerosa L, Hammel M, Bruning JM, Wendorff TJ, Phung W, Hymowitz SG, Sudhamsu J
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.2 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
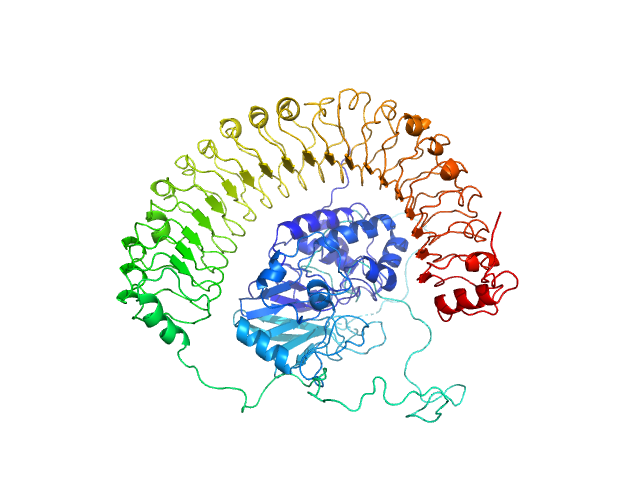
Leucine-rich repeat protein SHOC-2 monomer, 65 kDa Homo sapiens protein
Serine/threonine-protein phosphatase PP1-gamma catalytic subunit monomer, 37 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris pH 7.5, 100 mM NaCl, 1 mM TCEP, 1 mM MnCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Jul 20
|
Structural basis for SHOC2 modulation of RAS signalling.
Nature (2022)
Liau NPD, Johnson MC, Izadi S, Gerosa L, Hammel M, Bruning JM, Wendorff TJ, Phung W, Hymowitz SG, Sudhamsu J
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
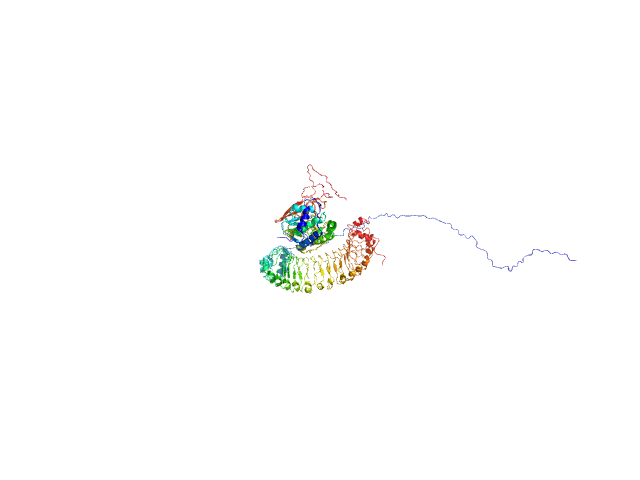
Leucine-rich repeat protein SHOC-2 monomer, 65 kDa Homo sapiens protein
Serine/threonine-protein phosphatase PP1-gamma catalytic subunit monomer, 37 kDa Homo sapiens protein
GTPase KRas monomer, 19 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris pH 7.5, 100 mM NaCl, 1 mM TCEP, 1 mM MnCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Jul 20
|
Structural basis for SHOC2 modulation of RAS signalling.
Nature (2022)
Liau NPD, Johnson MC, Izadi S, Gerosa L, Hammel M, Bruning JM, Wendorff TJ, Phung W, Hymowitz SG, Sudhamsu J
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
156 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
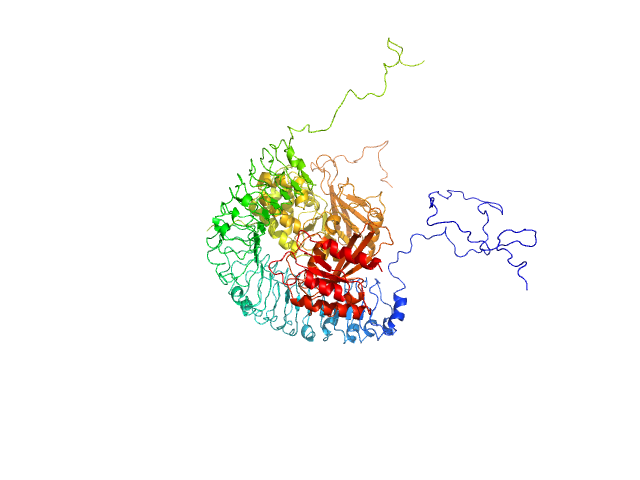
Leucine-rich repeat protein SHOC-2 monomer, 65 kDa Homo sapiens protein
Serine/threonine-protein phosphatase PP1-gamma catalytic subunit monomer, 37 kDa Homo sapiens protein
Ras-related protein M-Ras monomer, 24 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris pH 7.5, 100 mM NaCl, 1 mM TCEP, 1 mM MnCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Jul 20
|
Structural basis for SHOC2 modulation of RAS signalling.
Nature (2022)
Liau NPD, Johnson MC, Izadi S, Gerosa L, Hammel M, Bruning JM, Wendorff TJ, Phung W, Hymowitz SG, Sudhamsu J
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
215 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Accumulation associated protein monomer, 78 kDa Staphylococcus epidermidis (strain … protein
|
| Buffer: |
50 mM MOPS, 50 mM NaCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Oct 16
|
Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap.
J Mol Biol :167708 (2022)
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB
|
| RgGuinier |
14.7 |
nm |
| Dmax |
55.8 |
nm |
|
|
|
|
|
|
|
| Sample: |
Accumulation associated protein (mutant) monomer, 78 kDa Staphylococcus epidermidis (strain … protein
|
| Buffer: |
50 mM MOPS, 50 mM NaCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 1
|
Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap.
J Mol Biol :167708 (2022)
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB
|
| RgGuinier |
14.1 |
nm |
| Dmax |
57.2 |
nm |
|
|
|
|
|
|
|
| Sample: |
Accumulation associated protein (mutant) monomer, 78 kDa Staphylococcus epidermidis (strain … protein
|
| Buffer: |
50 mM MOPS, 50 mM NaCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 1
|
Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap.
J Mol Biol :167708 (2022)
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB
|
| RgGuinier |
14.4 |
nm |
| Dmax |
56.6 |
nm |
|
|
|
|
|
|
|
| Sample: |
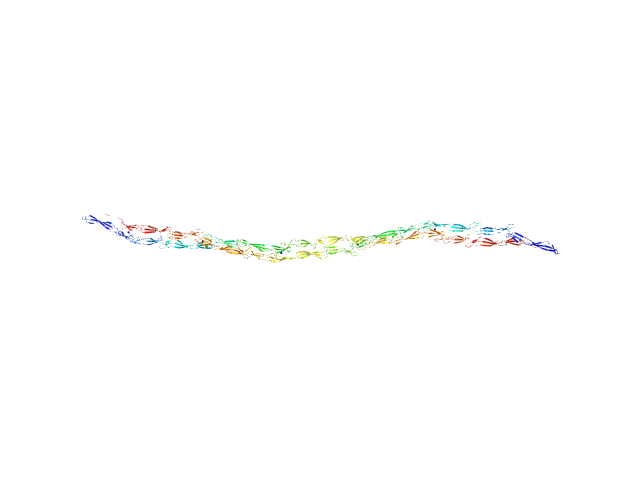
Accumulation associated protein (mutant) dimer, 155 kDa Staphylococcus epidermidis (strain … protein
|
| Buffer: |
50 mM MOPS, 50 mM NaCl, 5 mM ZnCl2, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 1
|
Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap.
J Mol Biol :167708 (2022)
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB
|
| RgGuinier |
15.0 |
nm |
| Dmax |
63.3 |
nm |
|
|
|
|
|
|
|
| Sample: |
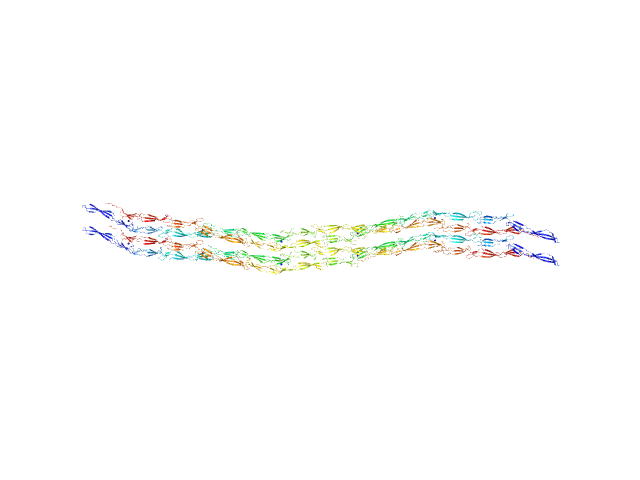
Accumulation associated protein (mutant) tetramer, 312 kDa Staphylococcus epidermidis (strain … protein
|
| Buffer: |
50 mM MOPS, 50 mM NaCl, 5 mM ZnCl2, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 1
|
Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap.
J Mol Biol :167708 (2022)
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB
|
| RgGuinier |
15.7 |
nm |
| Dmax |
70.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
Polyubiquitin-B monomer, 34 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02% NaN3, pH: 6.8
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Oct 8
|
Mechanistic insights into enhancement or inhibition of phase separation by different polyubiquitin chains.
EMBO Rep :e55056 (2022)
Dao TP, Yang Y, Presti MF, Cosgrove MS, Hopkins JB, Ma W, Loh SN, Castañeda CA
|
| RgGuinier |
2.6 |
nm |
| Dmax |
95.0 |
nm |
| VolumePorod |
44 |
nm3 |
|
|