|
X-ray synchrotron radiation scattering data from solutions of Human serum albumin in 50 mM HEPES, pH 7.5 were collected on the P12 beam line at Petra-III (Hamburg, Germany) using a Pilatus 2M detector (I(s) vs s, where s = 4π sin θ/λ; 2θ is the scattering angle and λ = 0.12 nm). SAXS data of the HSA monomer were obtained from averaging 38 x 1 second frames spanning a separated HSA monomer elution peak from in-line size exclusion chromatography-SAXS (top four profiles and the respective ab initio and atomic model fits). The bottom three profiles are derived from un-purified stock solutions of HSA, consisting of monomers, dimers, and higher oligomeric species at three concentrations: 20 mg/ml, 10 mg/ml and 5 mg/ml. These data are included to illustrate the effects of repulsive interparticle interactions. The atomic model monomer fits to the HSA mixtures (using PDB 1E78_chain_A) are also shown, with chi2 values of 1.99 (20 mg/ml), 1.27 (10 mg/ml) and 1.43 (5 mg/ml). The p-value determined using Correlation map for the monomer fit to the mixture data is 0 in all instances, indicating statistically significant differences are present between the model and data (cf. a chi2 of 1.01 and p-value of 0.035 for the purified monomer model fit to the SEC-SAXS data).
Cell temperature = UNKNOWN. X-ray Exposure time = UNKNOWN. Number of frames = UNKNOWN. Concentration = UNKNOWN
|
|
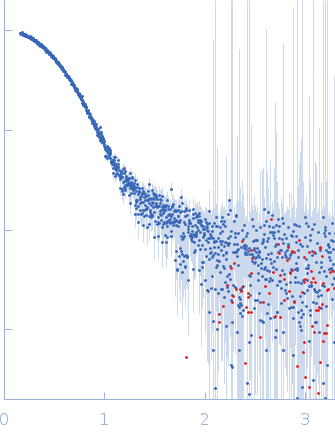 s, nm-1
s, nm-1