|
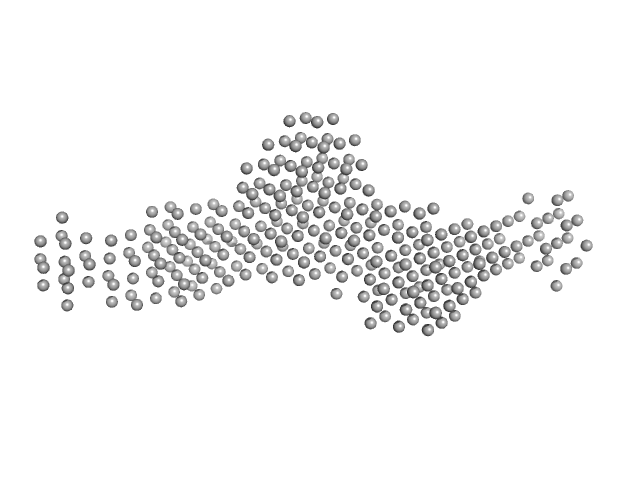
SAXS experiments were performed at the D02A-SAXS2 beamline at the Brazilian Synchrotron Light Laboratory (LNLS, Campinas, São Paulo, Brazil). The X-ray scattering data (I(s) vs s, where s = 4πsinθ/λ and 2θ is the scattering angle; λ = 0.1488 nm) were acquired using a two-dimensional position-sensitive MAR-CCD detector. Measurements were performed using a monochromatic X-ray beam and a sample-to-detector distance of ~1,000 mm, corresponding to the scattering vector range of 0.015 < q < 0.35 Å-1. LbSGT samples were prepared at 1.5 mg/mL, 3.0 mg/mL, 4.5 mg/mL and 6.0 mg/mL. Successive frames (20, 60 and 300 sec) were collected to inspect for X-ray damage. Aggregation or inter-particle interference were not observed. The model depicts the averaged spatial representation of the protein (DAMFILT occupancy and volume-corrected bead model).
|
|
SGT protein
(LbSGT)
|
| Mol. type |
|
Protein |
| Organism |
|
Leishmania braziliensis |
| Olig. state |
|
Dimer |
| Mon. MW |
|
45.9 kDa |
| |
| UniProt |
|
A4HIK6
|
| Sequence |
|
FASTA |
| |
|
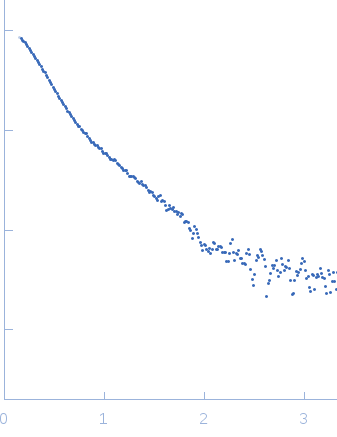 s, nm-1
s, nm-1
