|
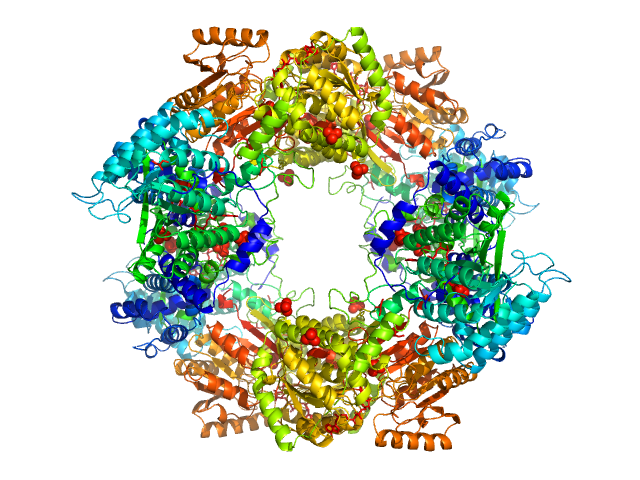
Synchrotron SAXS data from solutions of proline utilization A from Bradyrhizobium diazoefficiens (formerly Bradyrhizobium japonicum) were collected using SEC-SAXS in 50 mM Tris, 50 mM NaCl, 0.5 mM TCEP, 5% (v/v) glycerol, pH 7.8 on the BioCAT 18ID beam line at the Advanced Photon Source (APS, Argonne, IL, USA) using a Pilatus 100K detector at a sample-detector distance of 3.5 m and at a wavelength of λ = 0.103 nm (l(s) vs s, where s = 4πsinθ/λ and 2θ is the scattering angle). The SEC-SAXS elution was collected at 22°C. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
Storage temperature = UNKNOWN. X-ray Exposure time = UNKNOWN. Number of frames = UNKNOWN. SEC column = UNKNOWN. Sample injection volume = UNKNOWN. Flow rate = UNKNOWN
|
|
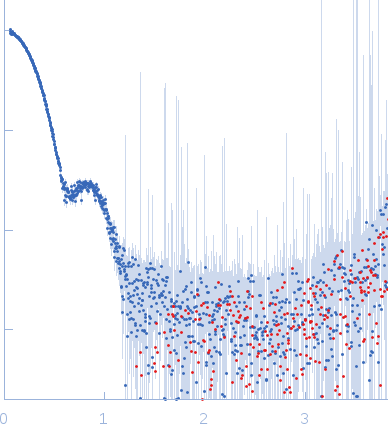 s, nm-1
s, nm-1