|
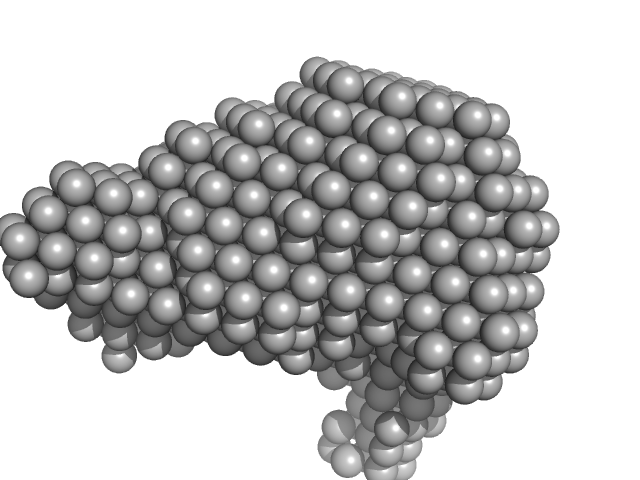
SAXS data from solutions of Ovalbumin monomer from in-house SEC-SAXS in PBS, pH 7.4 were collected on a Xenocs BioXolver L instrument at the Copenhagen University, Department of Drug Design and Pharmacology (Copenhagen, Denmark) using a 20Hz Pilatus 300K detector at a sample-detector distance of 0.7 m and at a wavelength of λ = 0.134 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
The OVA sample (500 microlitres at 4.9 mg/ml) was injected onto a GE Healthcare S200 Increase 10/300 (24 ml) column at a flow rate of 0.1 ml/min at 23°C. A total of 150 x 30 second SAXS data frames were recorded throughout the OVA elution. After background solvent corrections, SEC-SAXS data frames 90-100, corresponding to monomeric OVA, were scaled, averaged and binned logarithmically to produce the data displayed in this entry. Additional files, including plots of the selected frames and of I(0), HPLC and exposure cell UV traces and Rg vs. time (determined by AUTORG) are included in the additional files of the full entry zip archive.
|
|
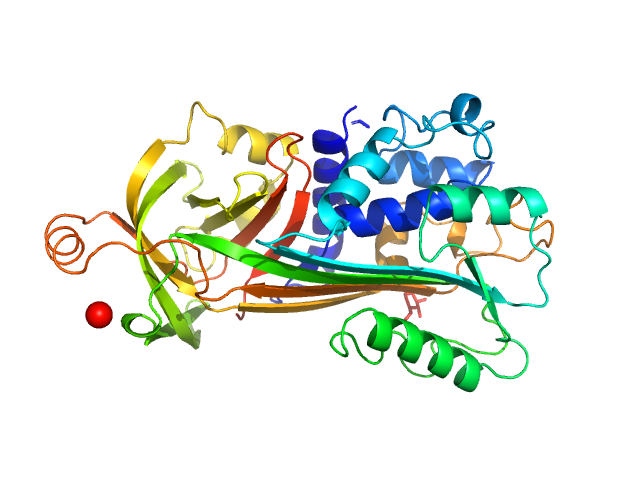
Ovalbumin
(OVA)
|
| Mol. type |
|
Protein |
| Organism |
|
Gallus gallus |
| Olig. state |
|
Monomer |
| Mon. MW |
|
42.8 kDa |
| |
| UniProt |
|
P01012
|
| Sequence |
|
FASTA |
| |
|
PDB ID
|
|
1OVA
|
| |
|
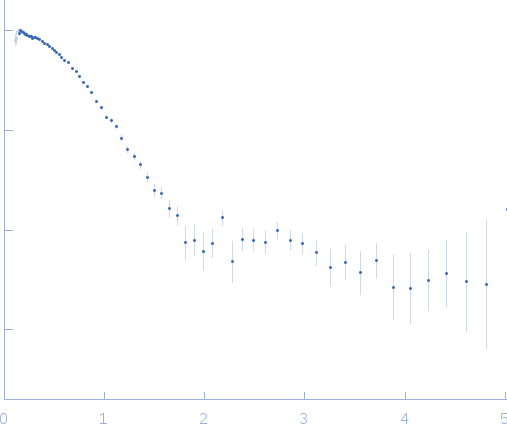 s, nm-1
s, nm-1