|
Synchrotron SAXS data from solutions of TraI from Neisseria gonorrhoeae in 50 mM TRIS-HCl, pH 8, 100 mM NaCl, were collected on the BM29 beam line at the ESRF storage ring (Grenoble, France) using a Pilatus 1M detector at a sample-detector distance of 2.9 m and at a wavelength of λ = 0.099 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Possible aggregates were removed using a Superdex 200 10/300 (GE Healthcare) size-exclusion chromatography (SEC) column pre-equilibrated with buffer at a flow rate of 0.5 mL/min at the EMBL-Lab outstation (Grenoble). All measurements were performed at 4°C. Sample concentration range was 0.7 to 10 mg/mL. For online-SEC one frame was collected every two seconds and for static measurements ten frames of data were collected with total exposure time of one second. By comparison of these frames, the possibility of radiation damage during the measurement was excluded. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. All used programs for data processing were part of the ATSAS Software package (Version 2.8.4).
|
|
TraI
(TraI)
|
| Mol. type |
|
Protein |
| Organism |
|
Neisseria gonorrhoeae |
| Olig. state |
|
Monomer |
| Mon. MW |
|
91.2 kDa |
| |
| UniProt |
|
Q5EPC7
(42-850)
|
| Sequence |
|
FASTA |
| |
|
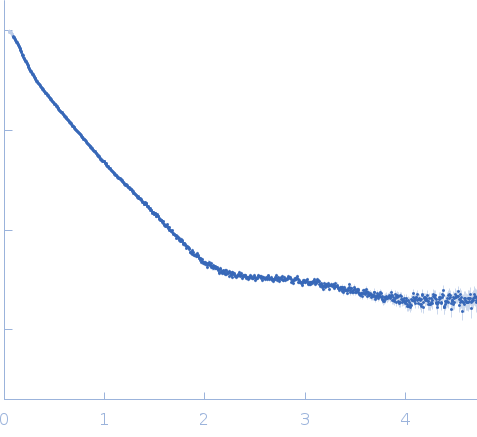 s, nm-1
s, nm-1
