|
SAXS data from solutions of the adenylation domain of DNA ligase A in 50 mM Tris-HCl 200 mM NaCl 2mM β-mercaptoethanol, pH 8 were collected on an Anton Paar SAXSpace instrument at the CSIR-Central Drug Research Institute (Lucknow, India) using a Hybrid Photon Counting (HPC) Mythen2 R 1K detector at a sample-detector distance of 0.3 m and at a wavelength of λ = 0.154 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). One solute concentration of 12.50 mg/ml was measured at 10°C. Two successive 1800 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
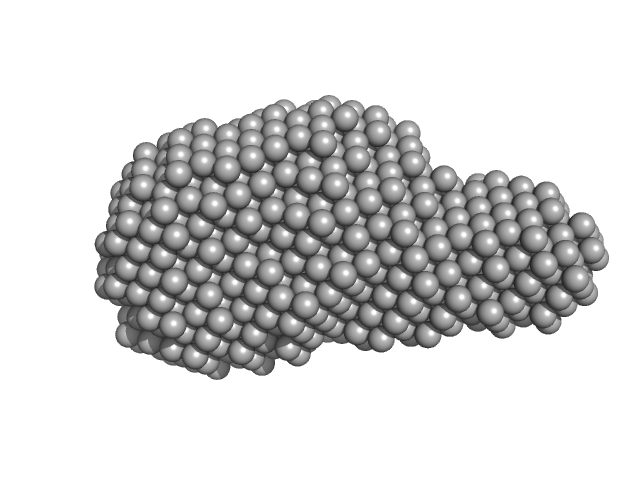
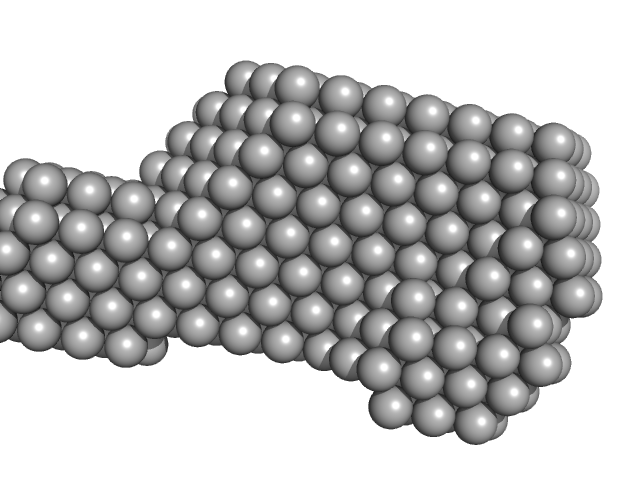
Ab initio dummy-atom models. Top = averaged spatial representation after multiple modelling runs (DAMFILT); Bottom = an example of an individual ab initio DAM model.
|
|
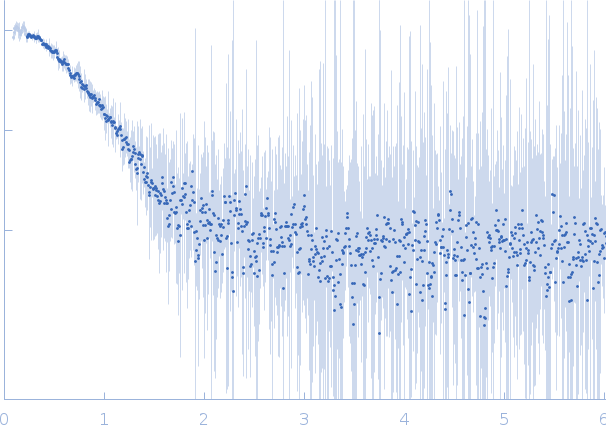 s, nm-1
s, nm-1