|
Synchrotron SAXS data from the 1:2 heterotrimer of pUL7 and pUL51(8-142) from herpes simplex virus 1 in 20 mM tris, 200 mM NaCl, 3% (v/v) glycerol, 0.25 mM TCEP, pH 7.5 were collected on the EMBL P12 beam line at PETRA III (Hamburg, Germany) using a Pilatus 6M detector at a sample-detector distance of 3 m and at a wavelength of λ = 0.124 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed. The profile displayed represents the average of three individual SEC-SAXS runs. The SEC parameters, per run, were as follows: A 75.00 μl sample at 4.5 mg/ml was injected at a 0.40 ml/min flow rate onto a GE Superdex 200 Increase 10/300 column at 20°C. Approximately 100 successive 1 second frames were collected through the major SEC-elution peak and processed using CHROMIXS. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
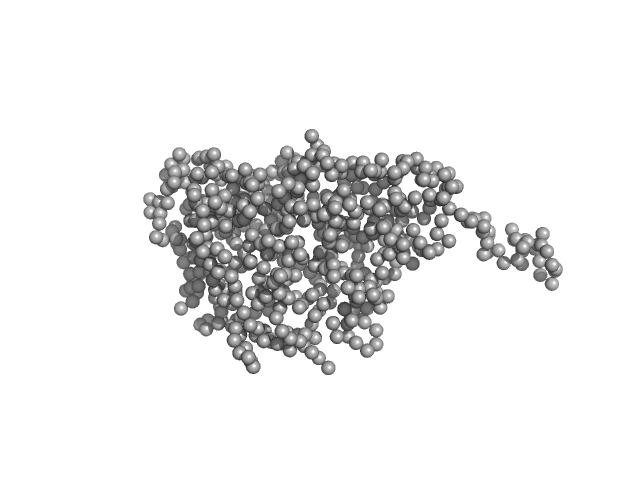
The quoted experimental molecular mass, that corresponds to a 1:2 pUL7:pUL51(8-142) heterotrimer, was determined from SEC-MALLS-RI measurements performed on identical samples, eluted in parallel to the SEC-SAXS. The SEC-SAXS data for each individual run, Rg correlations through the main elution peak as well as the the SEC-MALLS-RI data and hydrodynamic estimates obtained from QELS are made available in the full-entry zip archive. The CORAL rigid-body refined (atomistic) model displayed in this entry is one example of several alternatives whereby a second copy of pUL51 samples different spatial positions bound within the complex. Additional representations of the 1:2 pUL7:pUL51(8-142) heterotrimeric complex, including those incorporating mass spectrometry cross-linking distance constraints, are also made available in the full entry zip-archive.
|
|
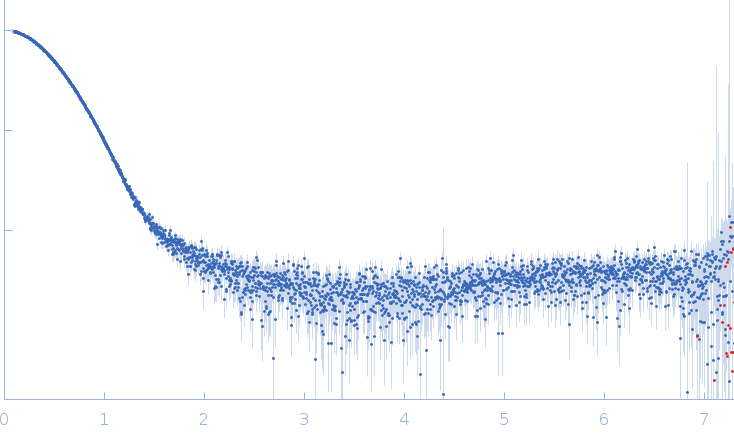 s, nm-1
s, nm-1