|
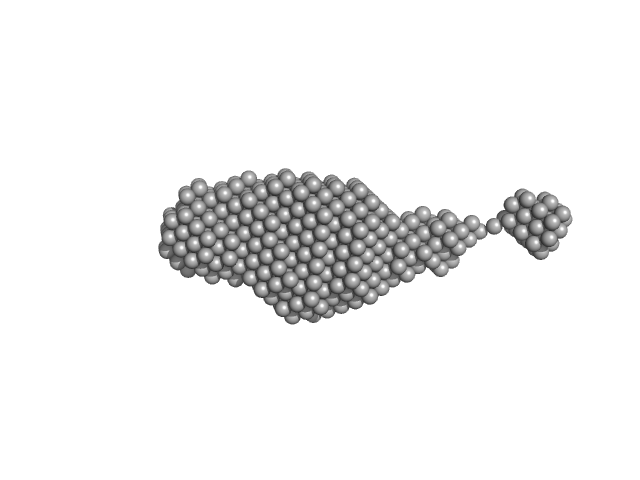
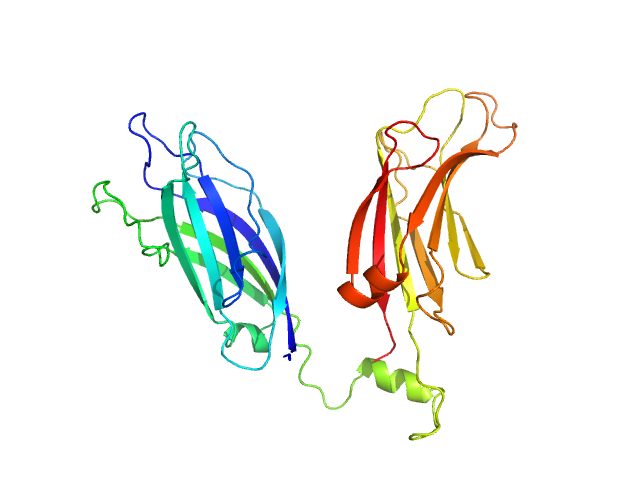
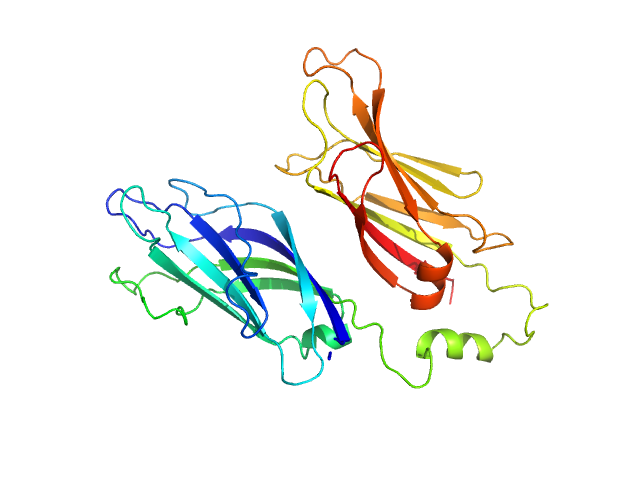
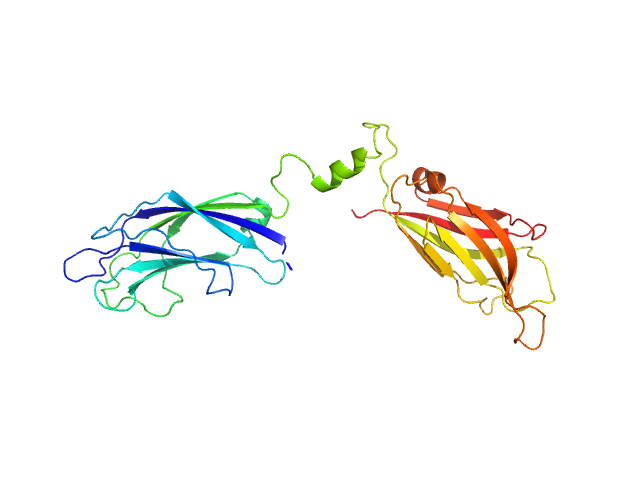
SAXS data were collected in the bioSAXS beamline B21, at Diamond Light Source, Harwell, United Kingdom. For the SYT1C2AB, 45 μl of sample with a concentration of 10 mg/ml, were injected onto a Superdex 200 increase 3.2/300 column at 20°C using 50 mM Tris, pH8, 50 mM NaCl as the running buffer. The output flow from the Agilent HPLC was directed through a 1.6 mm diameter quartz capillary cell held in vacuum. The flow rate was set to 0.08 mL min-1 and 1162 frames (with an exposure time of 3 s) were collected using a PILATUS 2M (Dectris, Switzerland) detector at the distance of 3.7 m from the sample. The collected two-dimensional images were corrected for variations in beam current, normalized for exposure time and processed into one- dimensional scattering curves using GDA and the DAWN software (Diamond Light Source, UK). The background was manually subtracted using the program ScÅtter and CHROMIXS (ATSAS).
Storage temperature = UNKNOWN
|
|
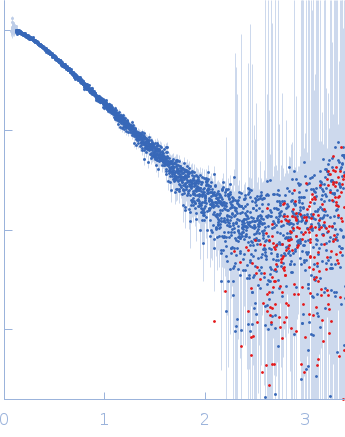 s, nm-1
s, nm-1