|
Synchrotron SAXS
data from solutions of
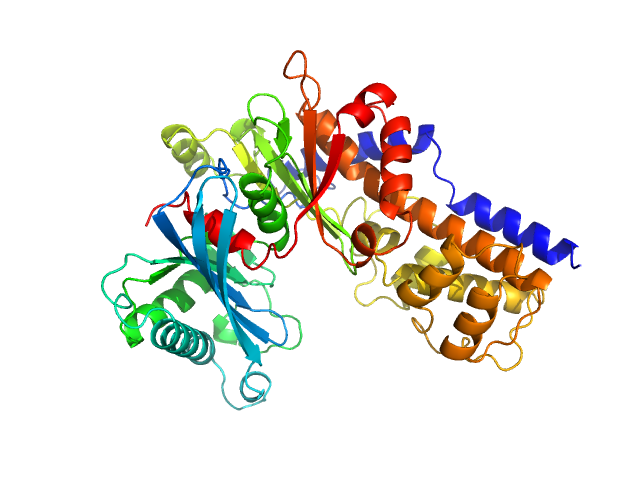
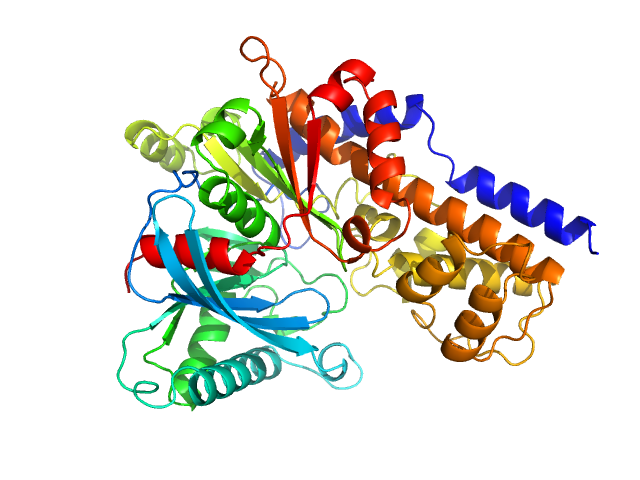
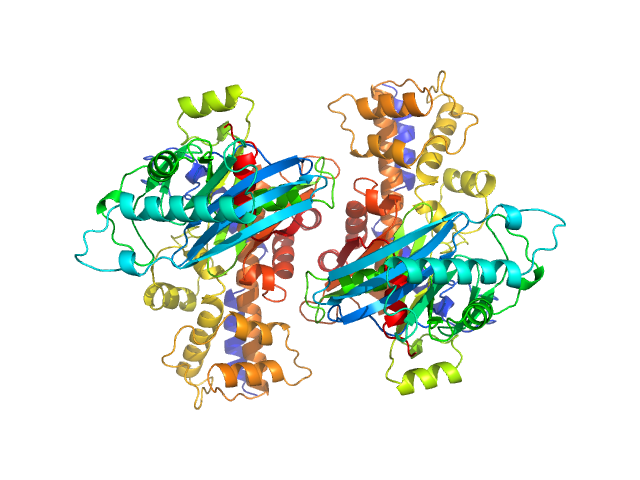
Monomeric Kluyveromyces lactis glucokinase 1 triple point mutant (I356D, Y419D, H420D), 3.0 mg/mL, no ligand
in
10 mM Tris-HCl, 1 mM DTT, pH 7.4
were collected
on the
EMBL P12 beam line
at the PETRA III storage ring
(DESY; Hamburg, Germany)
using a Pilatus 2M detector
at a sample-detector distance of 3 m and
at a wavelength of λ = 0.1239 nm
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
Solute concentrations ranging between 14.2 and 3 mg/ml were measured
at 10°C.
30 successive
0.045 second frames were collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
CAUTION: Repulsive interparticle interference effects present in the sample result in inaccurate structural parameters (Rg, I(0), MW, Dmax) extracted from the data.
|
|
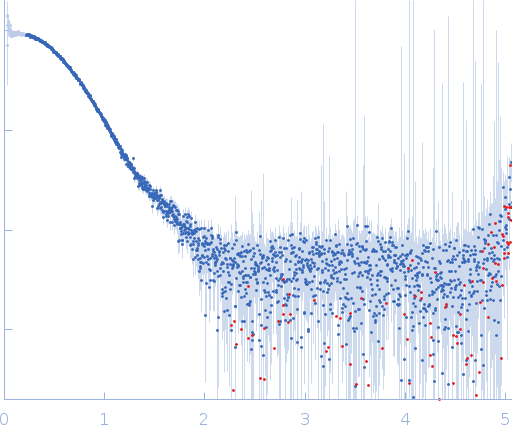 s, nm-1
s, nm-1