|
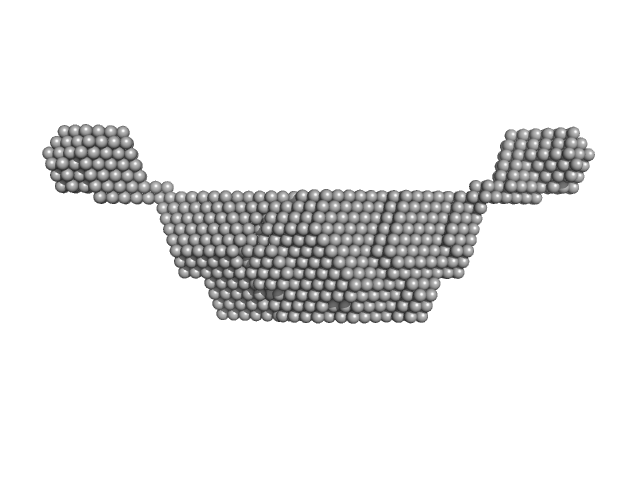
Synchrotron SAXS
data from solutions of
NTF2L domain and acidic IDR1 region of G3BP1 (NTF2LA, residues 1-232) at pH 6.0, 50 mM NaCl
in
50 mM MES, 50 mM NaCl,, pH 6
were collected
on the
B21 beam line
at the Diamond Light Source storage ring
(Didcot, UK)
using a Eiger 4M detector
at a sample-detector distance of 3.7 m and
at a wavelength of λ = 0.09464 nm
(I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle).
In-line size-exclusion chromatography (SEC) SAS was employed. The SEC parameters were as follows: A 60.00 μl sample
at 10 mg/ml was injected at a 0.08 ml/min flow rate
onto a Cytiva Superdex 75 Increase 3.2/300 column
at 15°C.
600 successive
3 second frames were collected.
The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
|
|
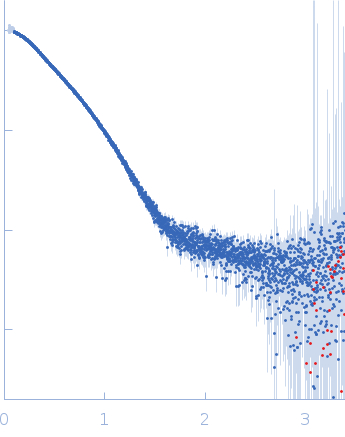 s, nm-1
s, nm-1
