|
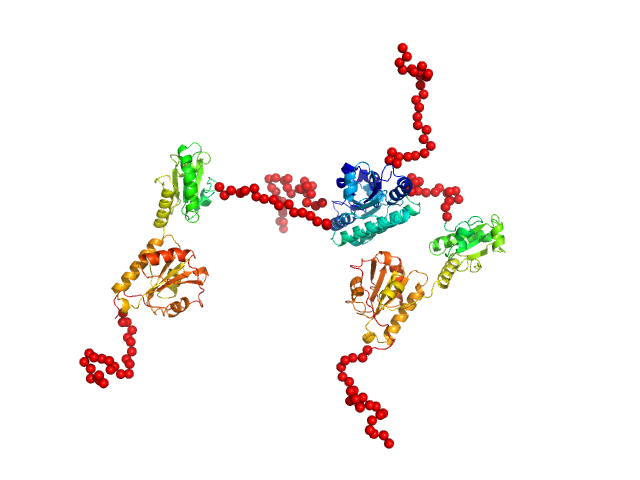
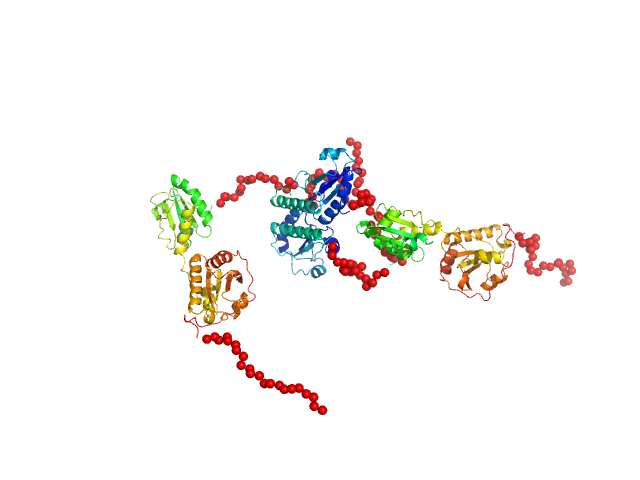
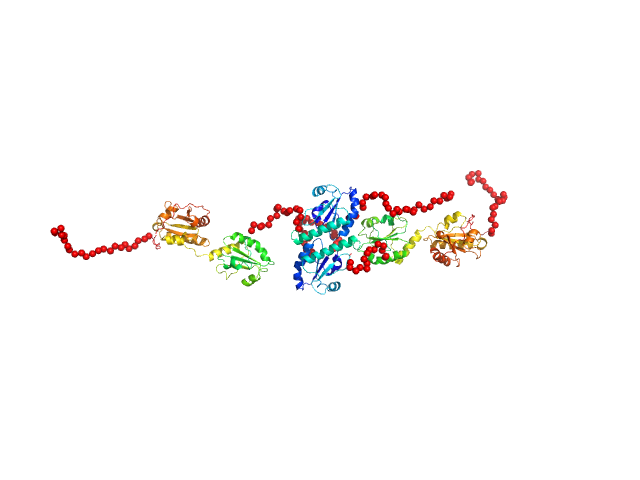
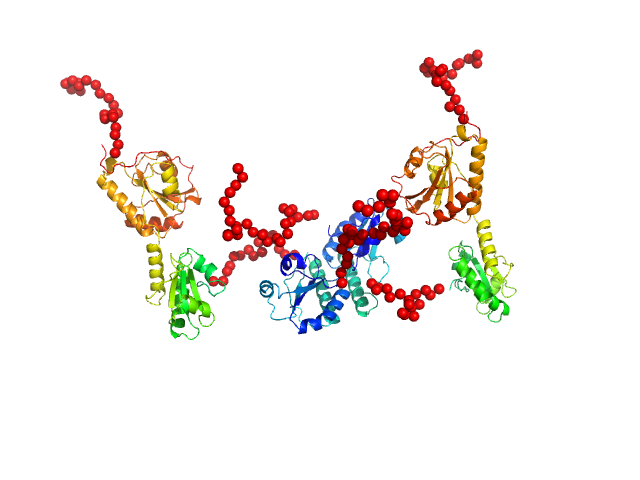
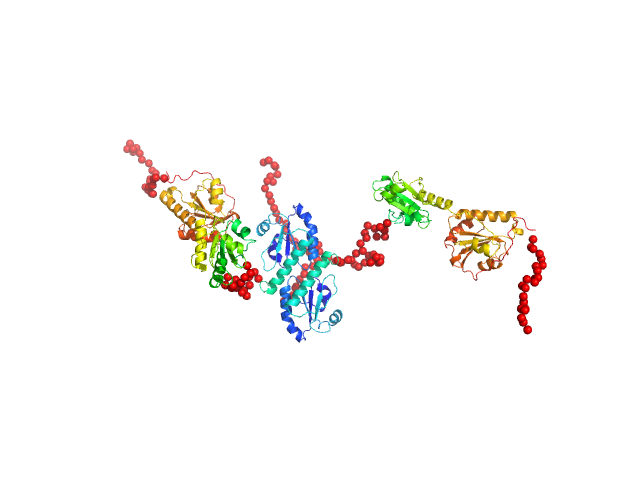
Synchrotron SEC–SAXS data from solutions of protein disulfide isomerase-like 2-3 precursor in 20 mM Tris-HCl, 200 mM NaCl, pH 8.0 were collected at beamline BL-15A2 of the Photon Factory (PF; Tsukuba, Ibaraki, Japan) using a PILATUS3 2M detector at a sample–detector distance of 2.65 m and a wavelength of λ = 0.10 nm (I(s) vs s, where s = 4π sin θ/λ and 2θ is the scattering angle). A total of 1,500 consecutive 3 s frames were recorded at 20 °C. The sample was loaded onto a Superdex 200 Increase 3.2/300 column (GE Healthcare) and eluted at 0.03 mL/min. The injected volume was 80 ul at a concentration of 3.6 mg/ml. Data were normalized to the incident beam intensity and azimuthally averaged; the solvent-blank scattering was subtracted. Frames within the region of constant Rg on the descending side of the elution peak (frames 915–960) were averaged using MOLASS. To estimate the molecular mass from I(0), the partial specific volume (0.74 cm^3/g) and the scattering contrast (2.798 × 10^10 cm^-2) were calculated using MULCh (http://smb-research.smb.usyd.edu.au/NCVWeb/input.jsp).
|
|
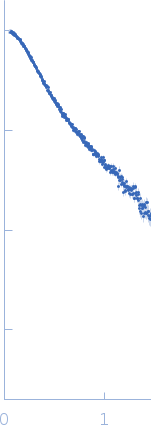 s, nm-1
s, nm-1
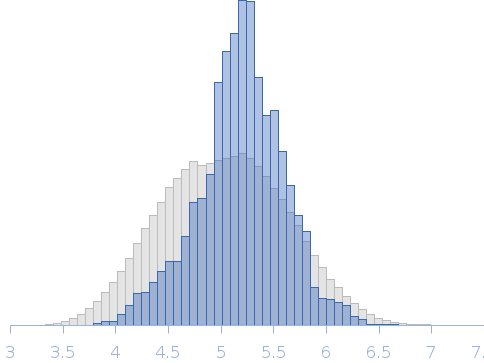 Rg, nm
Rg, nm