Structural basis for IL-33 recognition and its antagonism by the helminth effector protein HpARI2.
Jamwal A,
Colomb F,
McSorley HJ,
Higgins MK
Nat Commun
15(1):5226
(2024 Jun 19)
|
|
|
|

|
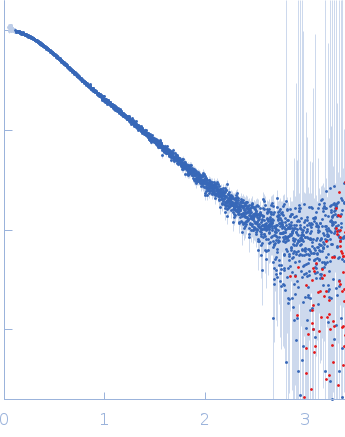
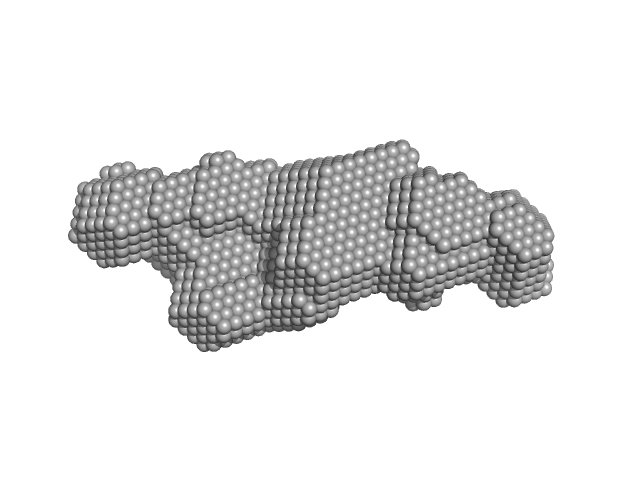
| Sample: |
Alarmin release inhibitor (Δ1-62) monomer, 26 kDa Heligmosomoides polygyrus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 5% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Feb 9
|
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
|
|
|
|

|
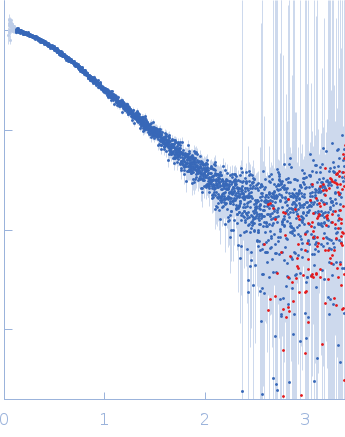
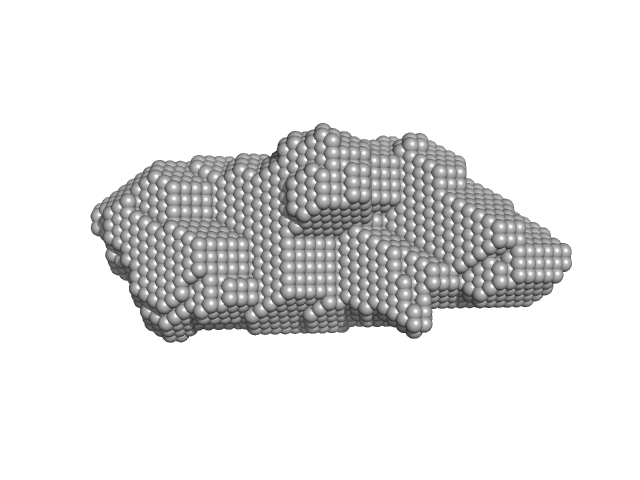
| Sample: |
Alarmin release inhibitor (Δ1-125; N175Q, N190Q) monomer, 19 kDa Heligmosomoides polygyrus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 5% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Feb 9
|
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|

|
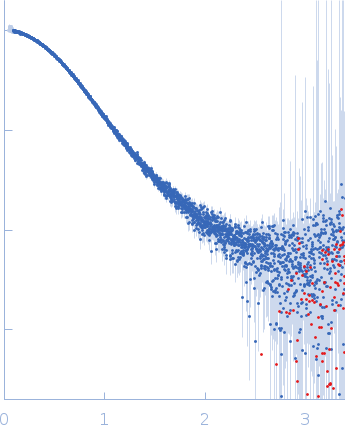
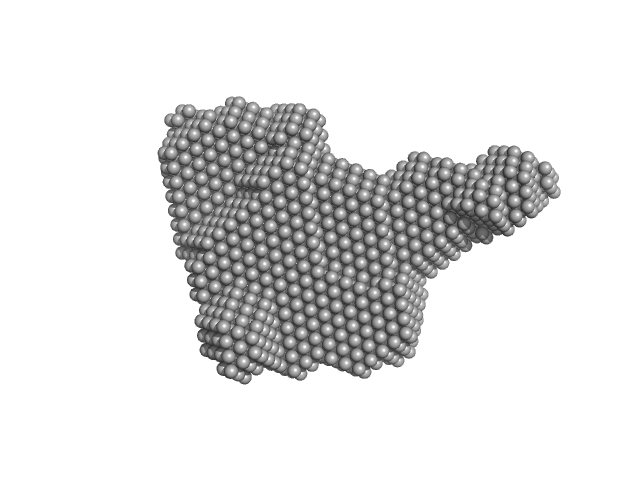
| Sample: |
Alarmin release inhibitor (Δ1-62) monomer, 26 kDa Heligmosomoides polygyrus protein
Interleukin-33 (L179V) monomer, 18 kDa Mus musculus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 5% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Feb 9
|
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
62 |
nm3 |
|
|
|
|
|
|
|
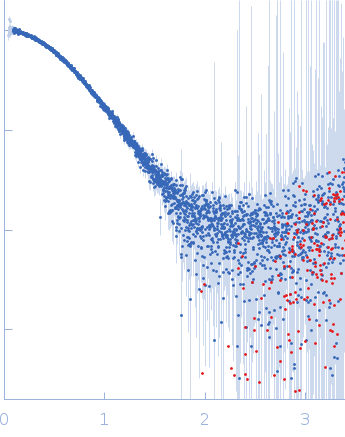
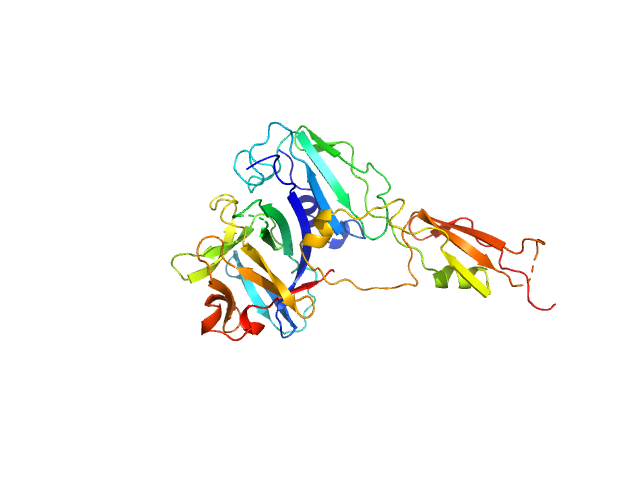
| Sample: |
Interleukin-33 (L179V) monomer, 18 kDa Mus musculus protein
Alarmin release inhibitor (Δ1-125; N175Q, N190Q) monomer, 19 kDa Heligmosomoides polygyrus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 5% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Feb 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
58 |
nm3 |
|
|