Allosteric activation of the nitric oxide receptor soluble guanylate cyclase mapped by cryo-electron microscopy.
Horst BG,
Yokom AL,
Rosenberg DJ,
Morris KL,
Hammel M,
Hurley JH,
Marletta MA
Elife
8
(2019 Sep 30)
|
|
|
|
|
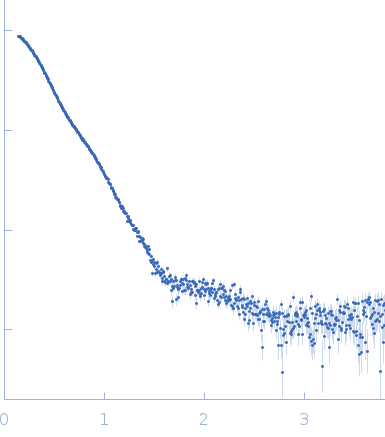
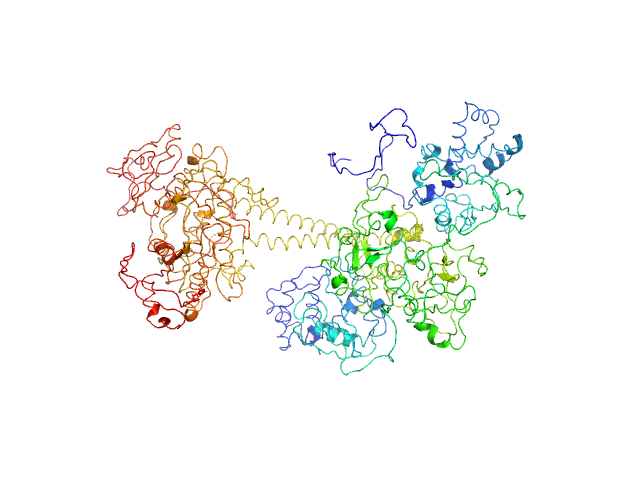
| Sample: |
Soluble guanylyl cyclase alpha-1 subunit monomer, 78 kDa Manduca sexta protein
Soluble guanylyl cyclase beta-1 subunit monomer, 68 kDa Manduca sexta protein
|
| Buffer: |
50 mM KH2PO4, 150 mM NaCl, 2% glycerol,, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Dec 3
|
|
| RgGuinier |
4.3 |
nm |
| Dmax |
13.3 |
nm |
| VolumePorod |
230 |
nm3 |
|
|
|
|
|
|
|
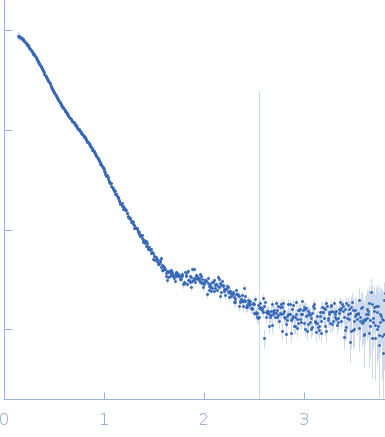
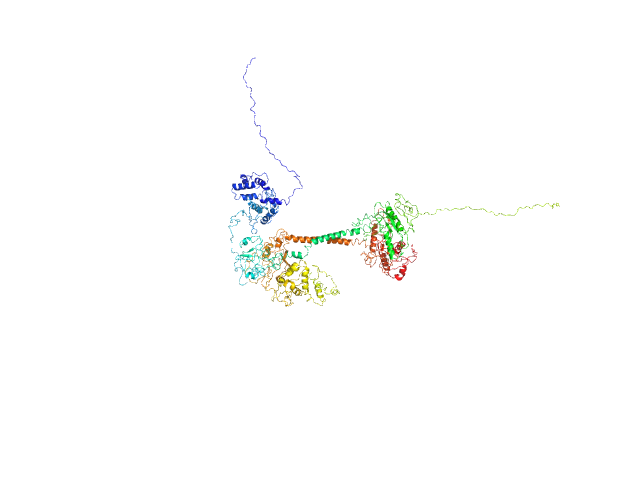
| Sample: |
Soluble guanylyl cyclase alpha-1 subunit monomer, 78 kDa Manduca sexta protein
Soluble guanylyl cyclase beta-1 subunit monomer, 68 kDa Manduca sexta protein
|
| Buffer: |
50 mM KH2PO4,150 mM NaCl, 2% glycerol, 500 µM NO,, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Dec 3
|
|
| RgGuinier |
4.4 |
nm |
| Dmax |
14.2 |
nm |
| VolumePorod |
218 |
nm3 |
|
|