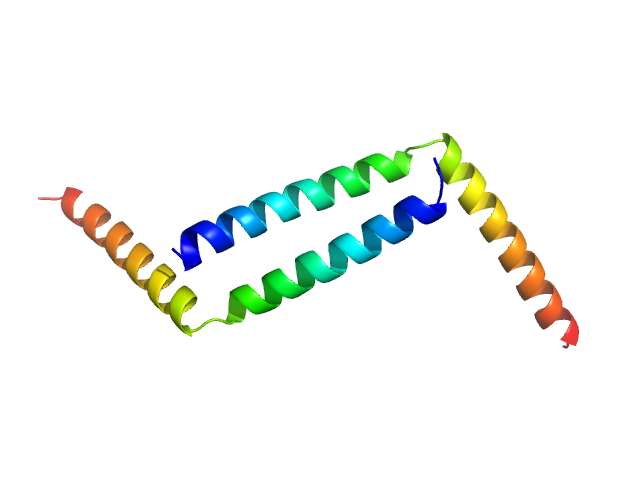|
Synchrotron SAXS data from solutions of TurB_nt50 in 20 mM Tris-HCl 500 mM NaCl, 10% glycerol, 500 mM Imidazole, pH8 were collected on the BL-10C beam line at the Photon Factory (PF), High Energy Accelerator Research Organization (KEK; Tsukuba, Japan) using a Pilatus3 2M detector at a sample-detector distance of 1.01 m and at a wavelength of λ = 0.15 nm (I(s) vs s, where s = 4πsinθ/λ and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed. The SEC parameters were as follows: A 80.00 μl sample at 14 mg/ml was injected at a 0.01 ml/min flow rate onto a GE Superdex 75 Increase 3.2/300 column at 20°C. 303 successive 20 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The averaged scattering profile near the elution peak (frames 134–144) was calculated by MOLASS.
|
|
H-NS family protein MvaT
(TurB_nt50)
|
| Mol. type |
|
Protein |
| Organism |
|
Pseudomonas putida (strain ATCC 47054 / DSM 6125 / CFBP 8728 / NCIMB 11950 / KT2440) |
| Olig. state |
|
Dimer |
| Mon. MW |
|
5.8 kDa |
| |
| UniProt |
|
Q88GF9
(1-50)
|
| Sequence |
|
FASTA |
| |
|
PDB ID
|
|
8H8H
|
| |
|
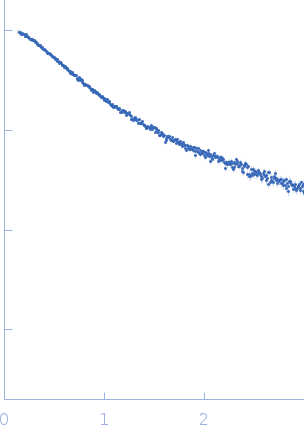 s, nm-1
s, nm-1