UniProt ID: P01308 (25-110) Human insulin
|
|
|

|
| Sample: |
Human insulin hexamer, 35 kDa Homo sapiens protein
|
| Buffer: |
7 mM sodium phosphate, 60 mM phenol, 200 µM Zn(CH3COO)2, 23 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, University of Copenhagen, Department of Drug Design and Pharmacology on 2018 Jan 26
|
Size-exclusion chromatography small-angle X-ray scattering of water soluble proteins on a laboratory instrument.
J Appl Crystallogr 51(Pt 6):1623-1632 (2018)
Bucciarelli S, Midtgaard SR, Nors Pedersen M, Skou S, Arleth L, Vestergaard B
|
| RgGuinier |
2.0 |
nm |
| Dmax |
5.9 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin (Insulin B chain and Insulin A chain)
UniProt ID: None (None-None) HUI-018 Fab
|
|
|
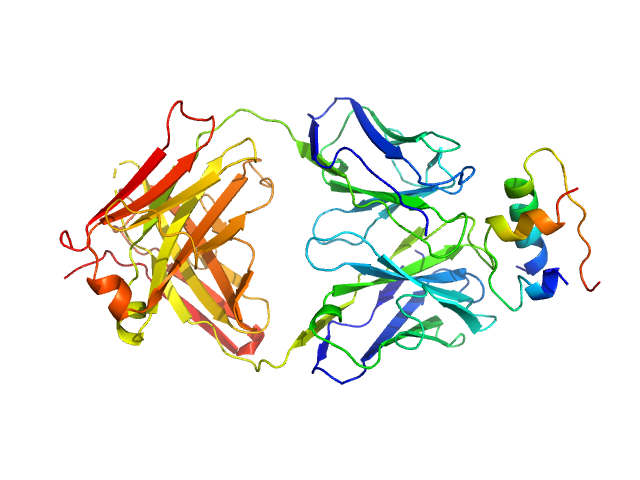
|
| Sample: |
Insulin (Insulin B chain and Insulin A chain) monomer, 6 kDa Homo sapiens protein
HUI-018 Fab monomer, 47 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2013 Mar 7
|
Insulin Binding to the Analytical Antibody Sandwich Pair OXI
‐005 and HUI
‐018 – Epitope Mapping and Binding Properties
Protein Science (2020)
Johansson E, Wu X, Yu B, Yang Z, Cao Z, Wiberg C, Jeppesen C, Poulsen F
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
86 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin (Insulin B chain and Insulin A chain)
UniProt ID: None (None-None) HUI-018 Fab
|
|
|
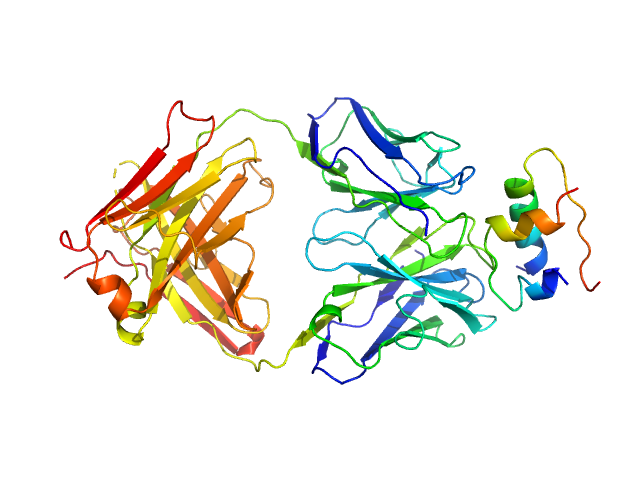
|
| Sample: |
Insulin (Insulin B chain and Insulin A chain) monomer, 6 kDa Homo sapiens protein
HUI-018 Fab monomer, 47 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2013 Mar 7
|
Insulin Binding to the Analytical Antibody Sandwich Pair OXI
‐005 and HUI
‐018 – Epitope Mapping and Binding Properties
Protein Science (2020)
Johansson E, Wu X, Yu B, Yang Z, Cao Z, Wiberg C, Jeppesen C, Poulsen F
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
105 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin detemir
|
|
|
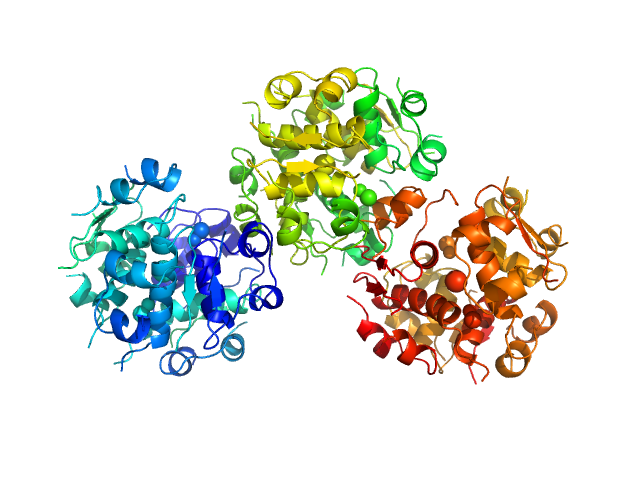
|
| Sample: |
Insulin detemir 18-mer, 106 kDa protein
|
| Buffer: |
5.0 mM Na2HPO4, 13.1 mM m-cresol, 15.1 mM phenol, 173.7 mM glycerol, 20.0 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.3 |
nm |
| VolumePorod |
137 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin detemir (Levemir(R), Novo Nordisk A/S)
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
|
|
|
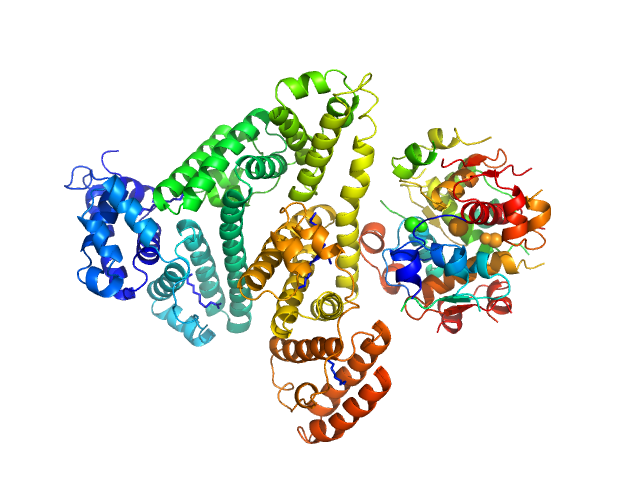
|
| Sample: |
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin detemir (Levemir(R), Novo Nordisk A/S)
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
|
|
|
|
| Sample: |
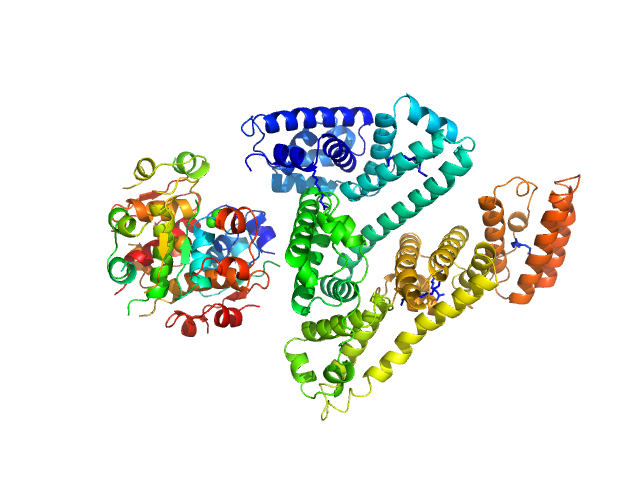
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
UniProt ID: P01308 (None-None) Insulin detemir (Levemir(R), Novo Nordisk A/S)
|
|
|
|
| Sample: |
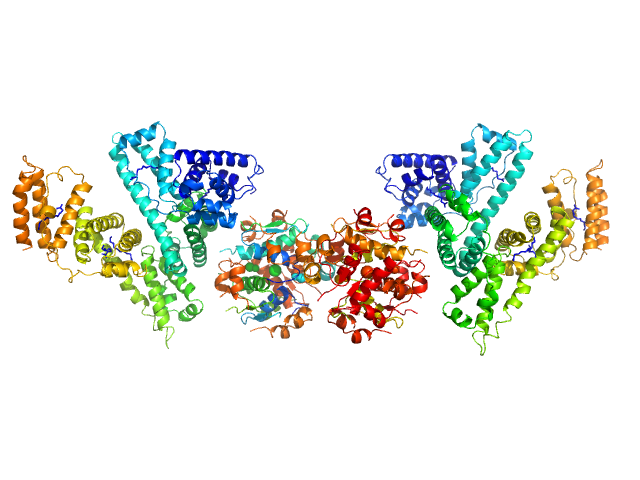
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
Insulin detemir (Levemir(R), Novo Nordisk A/S) dodecamer, 71 kDa protein
|
| Buffer: |
8.8 mM Na2HPO4, 10.6 mM m-cresol, 12.2 mM phenol, 140.9 mM glycerol, 56.9 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
309 |
nm3 |
|
|
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
UniProt ID: P01308 (None-None) Insulin detemir (Levemir(R), Novo Nordisk A/S)
|
|
|
|
| Sample: |
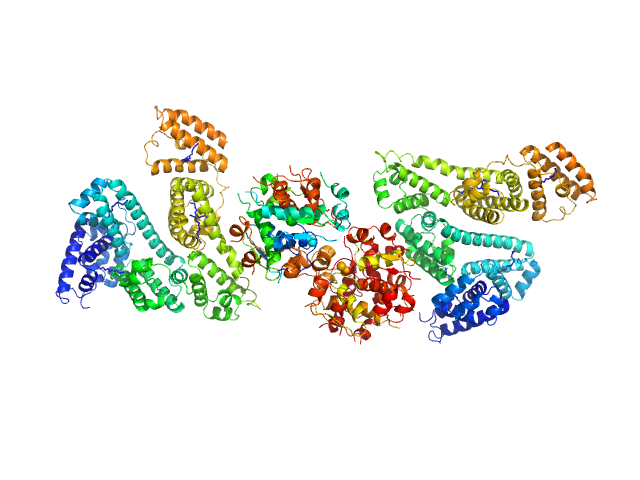
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
Insulin detemir (Levemir(R), Novo Nordisk A/S) dodecamer, 71 kDa protein
|
| Buffer: |
8.8 mM Na2HPO4, 10.6 mM m-cresol, 12.2 mM phenol, 140.9 mM glycerol, 56.9 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
309 |
nm3 |
|
|
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Elite, Albumedix Ltd.)
UniProt ID: P01308 (25-110) Insulin degludec(Tresiba(R), Novo Nordisk A/S)
|
|
|
|
| Sample: |
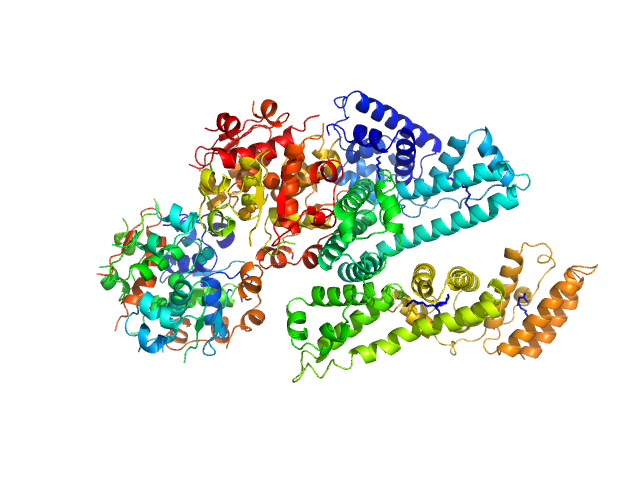
Human Albumin (Recombumin(R) Elite, Albumedix Ltd.) monomer, 66 kDa protein
Insulin degludec(Tresiba(R), Novo Nordisk A/S) dodecamer, 73 kDa protein
|
| Buffer: |
25 mM Na2HPO4, 15.9 mM m-cresol, 15.9 mM phenol, 212.8 mM glycerol, 20 mM NaCl, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Jun 27
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
179 |
nm3 |
|
|