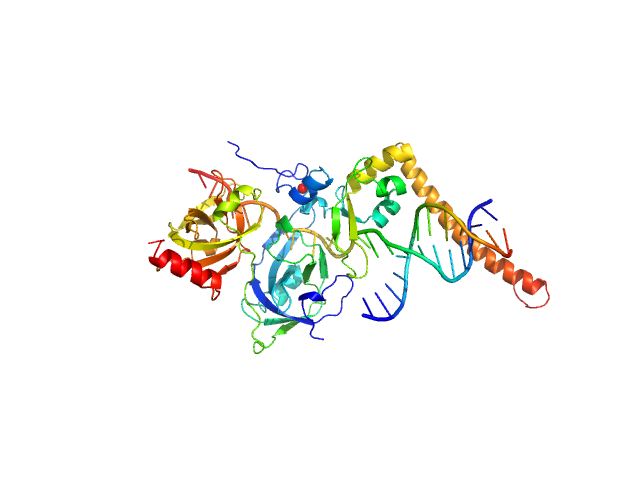
UniProt ID: P23025 (98-239) DNA repair protein complementing XP-A cells
UniProt ID: P27694 (183-420) Replication protein A 70 kDa DNA-binding subunit
UniProt ID: None (None-None) 3-prime Nucleotide Excision Repair Junction Model Substrate
|
|
|
|
| Sample: |
DNA repair protein complementing XP-A cells monomer, 17 kDa Homo sapiens protein
Replication protein A 70 kDa DNA-binding subunit monomer, 27 kDa Homo sapiens protein
3-prime Nucleotide Excision Repair Junction Model Substrate monomer, 11 kDa DNA
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 17 Nov 2
|
A key interaction with RPA orients XPA in NER complexes.
Nucleic Acids Res (2020)
Topolska-Woś AM, Sugitani N, Cordoba JJ, Le Meur KV, Le Meur RA, Kim HS, Yeo JE, Rosenberg D, Hammel M, Schärer OD, Chazin WJ
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
103 |
nm3 |
|
|
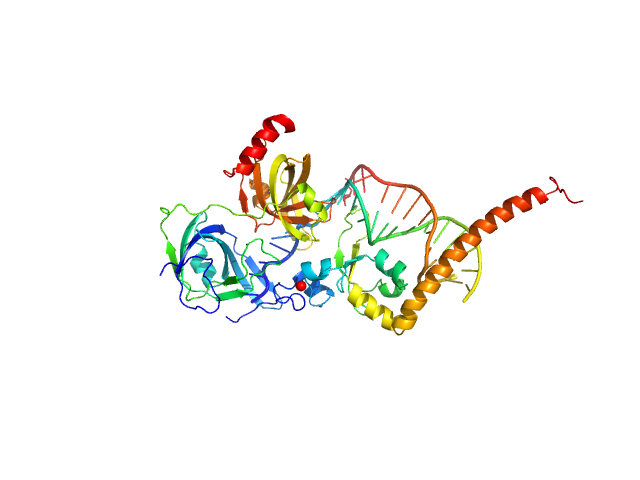
UniProt ID: P23025 (98-239) DNA repair protein complementing XP-A cells
UniProt ID: P27694 (183-420) Replication protein A 70 kDa DNA-binding subunit
UniProt ID: None (None-None) 5-prime Nucleotide Excision Repair Junction Model Substrate
|
|
|
|
| Sample: |
DNA repair protein complementing XP-A cells monomer, 17 kDa Homo sapiens protein
Replication protein A 70 kDa DNA-binding subunit monomer, 27 kDa Homo sapiens protein
5-prime Nucleotide Excision Repair Junction Model Substrate monomer, 11 kDa DNA
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Jun 4
|
A key interaction with RPA orients XPA in NER complexes.
Nucleic Acids Res (2020)
Topolska-Woś AM, Sugitani N, Cordoba JJ, Le Meur KV, Le Meur RA, Kim HS, Yeo JE, Rosenberg D, Hammel M, Schärer OD, Chazin WJ
|
| RgGuinier |
2.9 |
nm |
| Dmax |
97.0 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
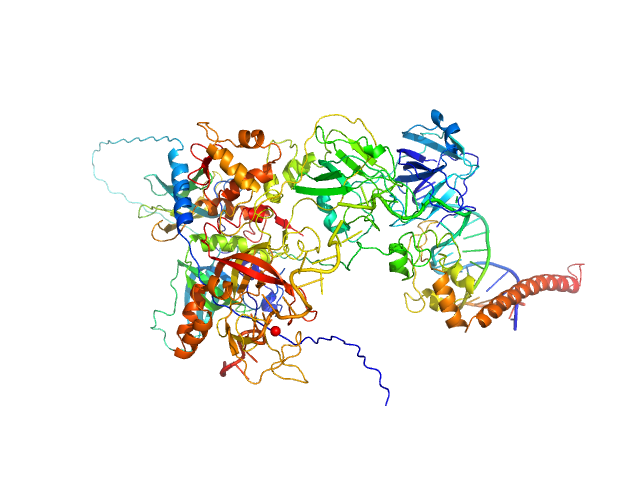
UniProt ID: P23025 (1-239) DNA repair protein complementing XP-A cells
UniProt ID: P27694 (185-616) Replication protein A 70 kDa DNA-binding subunit
UniProt ID: P15927 (45-270) Replication protein A 32 kDa subunit
UniProt ID: P35244 (1-121) Replication protein A 14 kDa subunit
UniProt ID: None (None-None) 3-prime ss-ds DNA junction NER model substrate
|
|
|
|
| Sample: |
DNA repair protein complementing XP-A cells monomer, 27 kDa Homo sapiens protein
Replication protein A 70 kDa DNA-binding subunit monomer, 49 kDa Homo sapiens protein
Replication protein A 32 kDa subunit monomer, 25 kDa Homo sapiens protein
Replication protein A 14 kDa subunit monomer, 14 kDa Homo sapiens protein
3-prime ss-ds DNA junction NER model substrate monomer, 17 kDa DNA
|
| Buffer: |
20 mM Tris pH 8.0, 150 mM NaCl, 2% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Mar 4
|
Two interaction surfaces between XPA and RPA organize the preincision complex in nucleotide excision repair
Proceedings of the National Academy of Sciences 119(34) (2022)
Kim M, Kim H, D’Souza A, Gallagher K, Jeong E, Topolska-Wós A, Ogorodnik Le Meur K, Tsai C, Tsai M, Kee M, Tainer J, Yeo J, Chazin W, Schärer O
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.7 |
nm |
| VolumePorod |
189 |
nm3 |
|
|
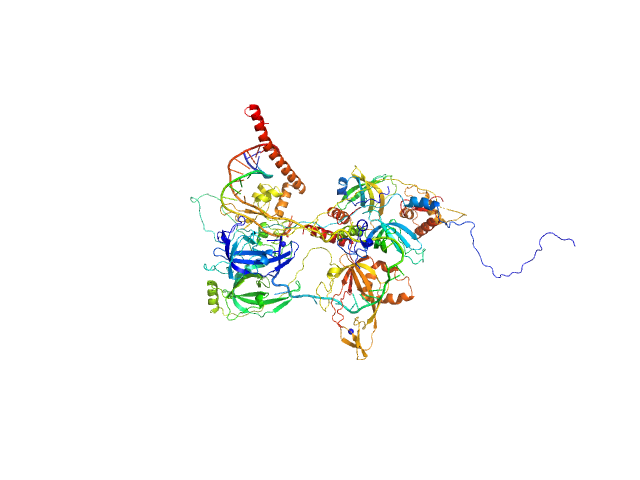
UniProt ID: P35244 (1-121) Replication protein A 14 kDa subunit
UniProt ID: P23025 (1-239) DNA repair protein complementing XP-A cells
UniProt ID: P27694 (185-616) Replication protein A 70 kDa DNA-binding subunit
UniProt ID: P15927 (45-270) Replication protein A 32 kDa subunit
UniProt ID: None (None-None) 5-prime ss-ds DNA junction NER model substrate
|
|
|
|
| Sample: |
Replication protein A 14 kDa subunit monomer, 14 kDa Homo sapiens protein
DNA repair protein complementing XP-A cells monomer, 27 kDa Homo sapiens protein
Replication protein A 70 kDa DNA-binding subunit monomer, 49 kDa Homo sapiens protein
Replication protein A 32 kDa subunit monomer, 25 kDa Homo sapiens protein
5-prime ss-ds DNA junction NER model substrate monomer, 17 kDa DNA
|
| Buffer: |
20 mM Tris pH 8.0, 150 mM NaCl, 2% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Mar 4
|
Two interaction surfaces between XPA and RPA organize the preincision complex in nucleotide excision repair
Proceedings of the National Academy of Sciences 119(34) (2022)
Kim M, Kim H, D’Souza A, Gallagher K, Jeong E, Topolska-Wós A, Ogorodnik Le Meur K, Tsai C, Tsai M, Kee M, Tainer J, Yeo J, Chazin W, Schärer O
|
| RgGuinier |
4.6 |
nm |
| Dmax |
16.5 |
nm |
| VolumePorod |
220 |
nm3 |
|
|