UniProt ID: P49419-2 (None-None) Aldehyde dehydrogenase 7A1 (Alpha-aminoadipic semialdehyde dehydrogenase)
|
|
|
|
| Sample: |
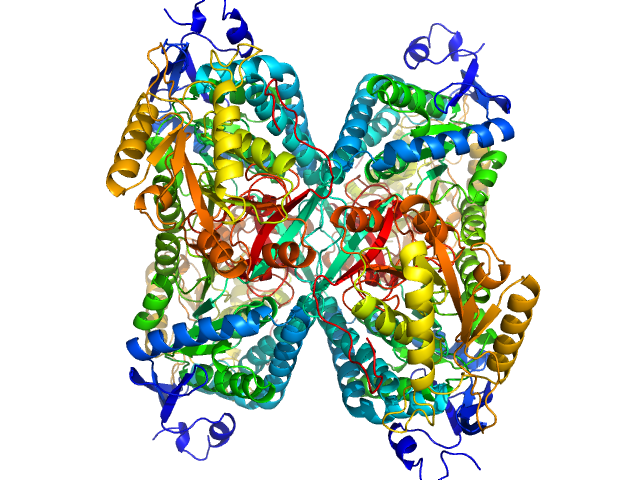
Aldehyde dehydrogenase 7A1 (Alpha-aminoadipic semialdehyde dehydrogenase) tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 5% glycerol, 0.5 mM tris(3-hydroxypropyl)phosphine, 50 mM NaCl, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Mar 9
|
Structural Basis of Substrate Recognition by Aldehyde Dehydrogenase 7A1.
Biochemistry 54(35):5513-22 (2015)
Luo M, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
270 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
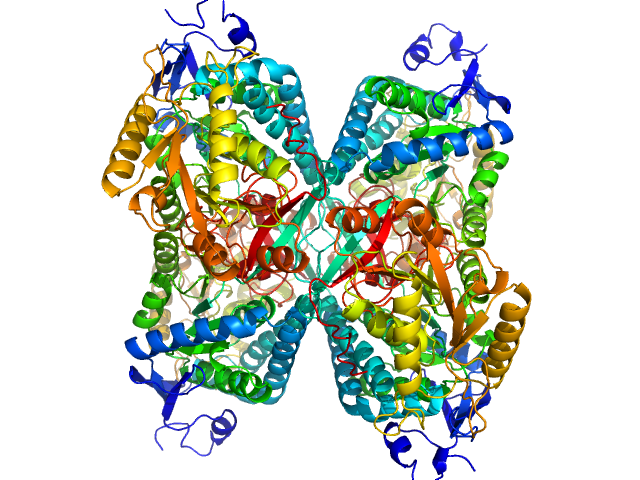
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.5 |
nm |
| VolumePorod |
212 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
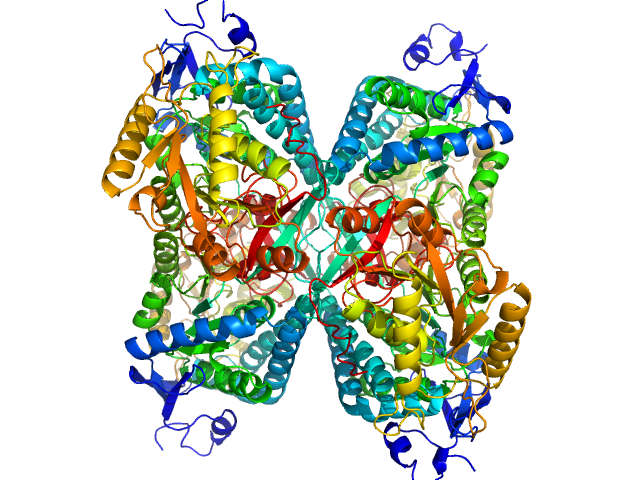
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.7 |
nm |
| VolumePorod |
229 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
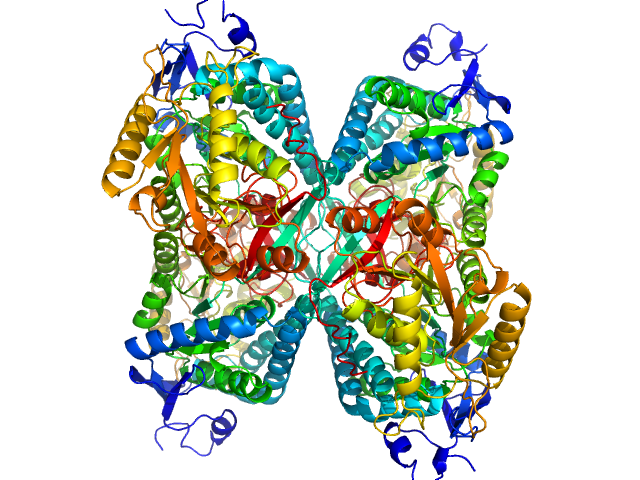
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.7 |
nm |
| VolumePorod |
238 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
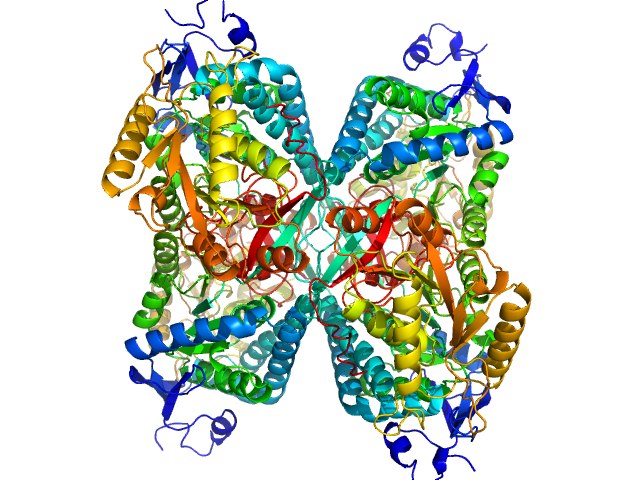
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, 1 mM NAD, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.7 |
nm |
| VolumePorod |
277 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
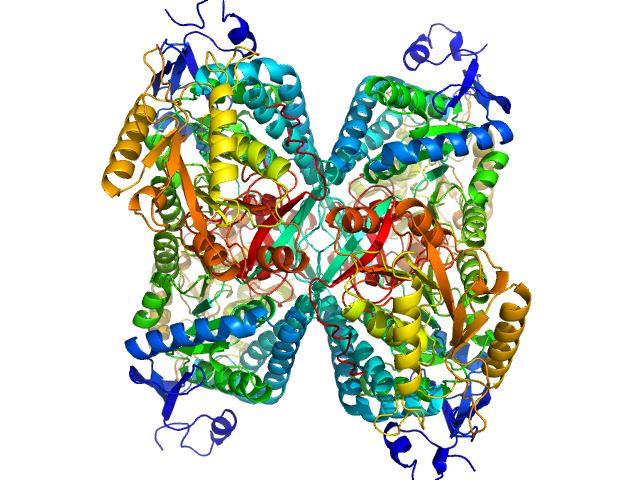
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, 1 mM NAD, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| VolumePorod |
275 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
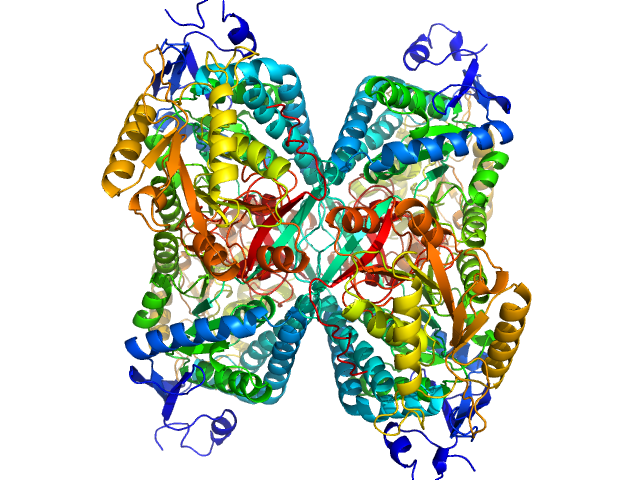
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 0.5 mM DTT, 5% (v/v) glycerol, 1 mM NAD, pH: 7.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Feb 22
|
NAD+ Promotes Assembly of the Active Tetramer of Aldehyde Dehydrogenase 7A1.
FEBS Lett (2018)
Korasick DA, White TA, Chakravarthy S, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| VolumePorod |
277 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
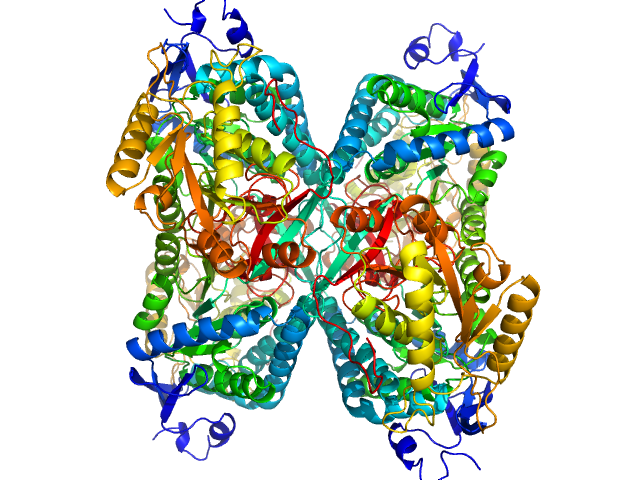
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 1 mM DTT, 10 mM NAD, 2% (v/v) glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 1
|
Structural Analysis of Pathogenic Mutations Targeting Glu427 of ALDH7A1, the Hot Spot Residue of Pyridoxine-Dependent Epilepsy.
J Inherit Metab Dis (2019)
Laciak AR, Korasick DA, Gates KS, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
350 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
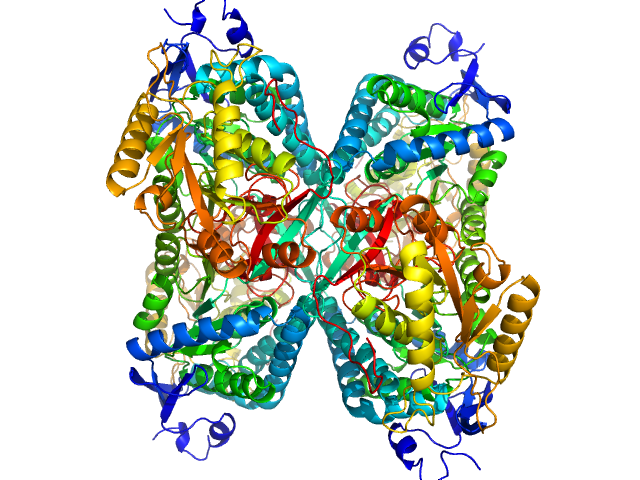
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 1 mM DTT, 10 mM NAD, 2% (v/v) glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 1
|
Structural Analysis of Pathogenic Mutations Targeting Glu427 of ALDH7A1, the Hot Spot Residue of Pyridoxine-Dependent Epilepsy.
J Inherit Metab Dis (2019)
Laciak AR, Korasick DA, Gates KS, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
326 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
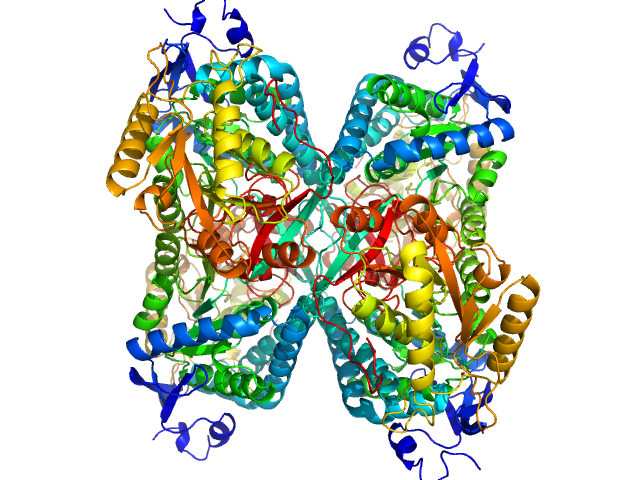
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 1 mM DTT, 10 mM NAD, 2% (v/v) glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 1
|
Structural Analysis of Pathogenic Mutations Targeting Glu427 of ALDH7A1, the Hot Spot Residue of Pyridoxine-Dependent Epilepsy.
J Inherit Metab Dis (2019)
Laciak AR, Korasick DA, Gates KS, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
315 |
nm3 |
|
|