|
|
|
|
|
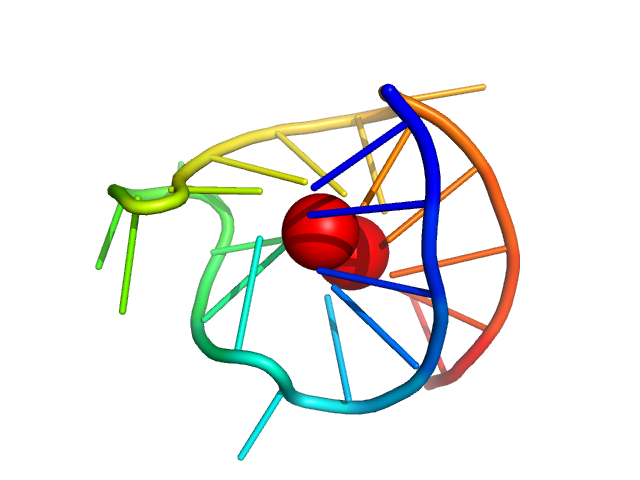
| Sample: |
VEGF promoter GQ parallel monomer, 7 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Nov 20
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.3 |
nm |
| Dmax |
3.8 |
nm |
| VolumePorod |
8 |
nm3 |
|
|
|
|
|
|
|
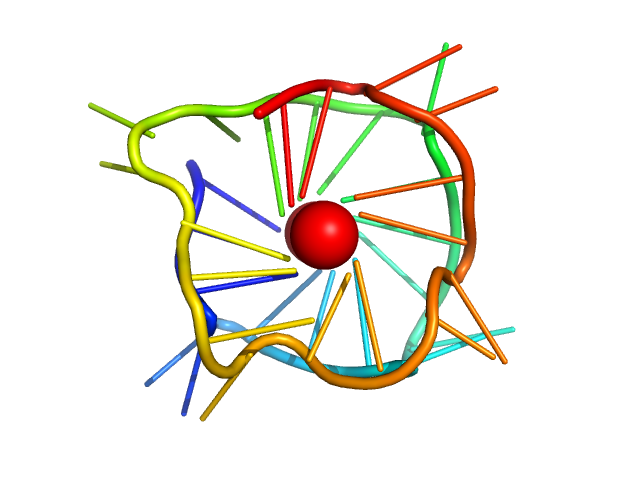
| Sample: |
cMyc promoter 8-tract G-quadruplex monomer, 11 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Feb 19
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.6 |
nm |
| VolumePorod |
13 |
nm3 |
|
|
|
|
|
|
|
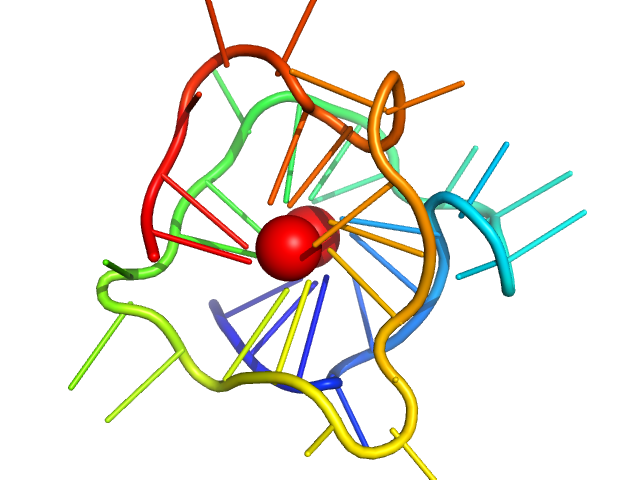
| Sample: |
cKit promoter 8-tract G-quadruplex monomer, 12 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Feb 19
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.7 |
nm |
| VolumePorod |
15 |
nm3 |
|
|
|
|
|
|
|
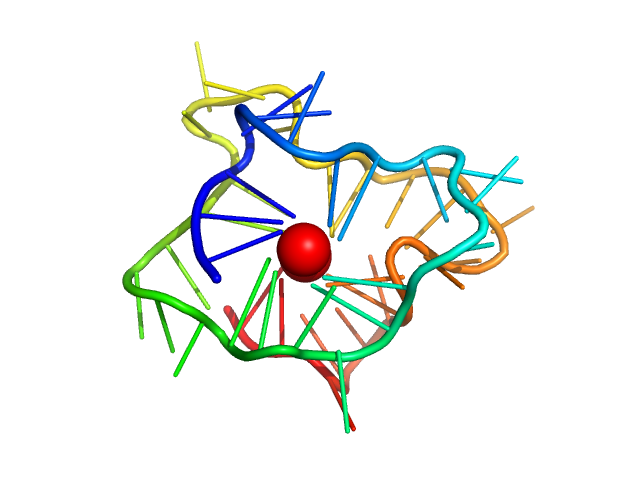
| Sample: |
kRas promoter 8-tract G-quadruplex monomer, 14 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Feb 19
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.7 |
nm |
| Dmax |
5.6 |
nm |
| VolumePorod |
16 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
cMyc promoter 12-tract G-quadruplex monomer, 23 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 22
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
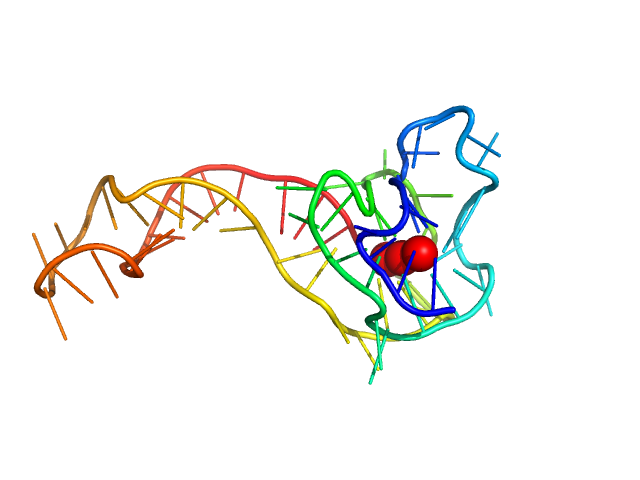
| Sample: |
cKit promoter 12-tract G-quadruplex monomer, 20 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Mar 22
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|

|
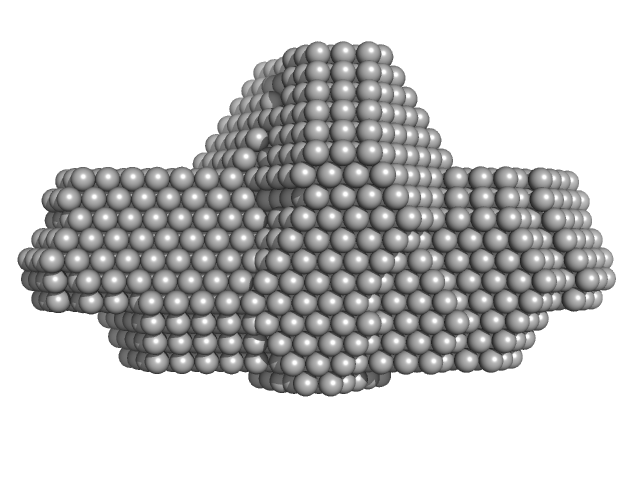
| Sample: |
Type I restriction-modification system methyltransferase subunit dimer, 144 kDa Vibrio vulnificus (strain … protein
Protein Ocr dimer, 28 kDa Escherichia phage T7 protein
|
| Buffer: |
20 mM Tris-HCl,150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2020 Apr 22
|
Structural features of a minimal intact methyltransferase of a type I restriction-modification system.
Int J Biol Macromol (2022)
Seo PW, Hofmann A, Kim JH, Hwangbo SA, Kim JH, Kim JW, Huynh TYL, Choy HE, Kim SJ, Lee J, Lee JO, Jin KS, Park SY, Kim JS
|
| RgGuinier |
4.2 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
354 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Transcription initiation factor II D monomer, 31 kDa Escherichia coli protein
|
| Buffer: |
50 mM HEPES, 5% v/v ethylene glycol, 2.5% v/v DMSO and 2 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Oct 26
|
Discovery of Dual TAF1-ATR Inhibitors and Ligand-Induced Structural Changes of the TAF1 Tandem Bromodomain.
J Med Chem 65(5):4182-4200 (2022)
Karim RM, Yang L, Chen L, Bikowitz MJ, Lu J, Grassie D, Shultz ZP, Lopchuk JM, Chen J, Schönbrunn E
|
| RgGuinier |
2.6 |
nm |
| Dmax |
7.3 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Transcription initiation factor II D monomer, 31 kDa Escherichia coli protein
|
| Buffer: |
50 mM HEPES, 5% v/v ethylene glycol, 2.5% v/v DMSO and 2 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Oct 19
|
Discovery of Dual TAF1-ATR Inhibitors and Ligand-Induced Structural Changes of the TAF1 Tandem Bromodomain.
J Med Chem 65(5):4182-4200 (2022)
Karim RM, Yang L, Chen L, Bikowitz MJ, Lu J, Grassie D, Shultz ZP, Lopchuk JM, Chen J, Schönbrunn E
|
| RgGuinier |
2.3 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Transcription initiation factor II D monomer, 31 kDa Escherichia coli protein
|
| Buffer: |
50 mM HEPES, 5% v/v ethylene glycol, 2.5% v/v DMSO and 2 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Oct 19
|
Discovery of Dual TAF1-ATR Inhibitors and Ligand-Induced Structural Changes of the TAF1 Tandem Bromodomain.
J Med Chem 65(5):4182-4200 (2022)
Karim RM, Yang L, Chen L, Bikowitz MJ, Lu J, Grassie D, Shultz ZP, Lopchuk JM, Chen J, Schönbrunn E
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
106 |
nm3 |
|
|